Table S1. Phosphopeptides regulated by SSE and insulin.
|
|
|
- Gunhild Olafsen
- 4 år siden
- Visninger:
Transkript
1 Table S. Phosphopeptides regulated by SSE and insulin. Phospho Site AAADDG EEPKS(ph) EPETK AAADDG EEPKS(ph) EPETKK AEDGAA PS(ph)PSSE T(ph)PKKK AEDGAA PSPSSET(p h)pkk AEGSLHS S(ph)PAGP SSSK AENQRP AEDSALS( ph)pgpla GAK AHTDEP HS(ph)PQS KK ALRPS(ph )HR ALSRS(ph )NEQMVS EKPSESK APLKDE QEMRAS(p h)pk AQGKPV SGQESSQS (ph)pyer AQQPAP ASS(ph)PV KR ARNDES QLS(ph)PA TR AS(ph)PG GVSTSSSD GKAEK ATAPQT QHVS(ph)P MR AYRPPS(p h)gek DDTSRYD ERPGPS(ph )PLPHR DGLPRTP S(ph)R DKPRS(ph )PFSK EAGS(ph) EAESSSAD PGPGRK Entrez Gene Name ID Phospho Fold Change apurinic/apyrimidinic endodeoxyribonuclease apurinic/apyrimidinic endodeoxyribonuclease myristoylated alanine rich protein kinase C substrate myristoylated alanine rich protein kinase C substrate zinc finger CCCH-type containing 4 LRR binding FLII interacting protein p-value Regulation by SSE Regulation by insulin P E-07 upregulation upregulation P E-09 upregulation upregulation P E-05 upregulation upregulation P E-06 upregulation upregulation Q6ZPZ E-07 upregulation upregulation Q3UZ E-06 upregulation upregulation elac ribonuclease Z 2 Q80Y E-05 upregulation upregulation SUMO/sentrin/SMT3 Q9EP E-06 upregulation upregulation specific peptidase 3 calpastatin P upregulation upregulation MYC binding protein 2, E3 ubiquitin protein ligase Q7TPH E-08 upregulation upregulation SEC3 homolog A, Q3UPL E-0 upregulation upregulation COPII coat complex component megakaryoblastic Q8K4J E-07 upregulation upregulation leukemia (translocation) plectin Q9QXS E-0 upregulation upregulation atrophin O E-07 upregulation upregulation eukaryotic translation elongation factor delta natural killer cell triggering receptor small nuclear ribonucleoprotein U subunit 70 RAB family interacting protein 5 ribosomal RNA processing homolog (S. cerevisiae) P upregulation upregulation P E-07 upregulation upregulation Q upregulation upregulation Q8BTI E-07 upregulation upregulation A0A0N4S E-06 upregulation upregulation W73 P E-07 upregulation upregulation
2 EFKENKE PS(ph)PK EFTGS(ph )PPPSATK K EGLS(ph) PAKR EIKPSEKP VS(ph)PK EKENGFS S(ph)PPR ERSVS(ph )VDSGEQR ERTSS(ph) LTHSEEK ESS(ph)PS HGLLK ETNVSKE DT(ph)DQ EEKASNE DVTK FGLHGS( ph)sgk FT(ph)DT RKDEQER GGDGAP RGS(ph)PS PASVSSGR GGGQSS( ph)pqeept WKK GGLPVPK KS(ph)PK GGSSEEL HDS(ph)PR GINGGPS RMS(ph)PK GINGGPS RMS(ph)PK GIS(ph)SS NEGVEEPS K GKLQEE GGS(ph)EE EEAGNPSE DGMQSGP TQAPPR GRAS(ph) PGGVSTSS SDGK GREEDSS( ph)pgpqs QSHRT GRTSST(p h)nededl NPEQK myosin IXA Q8C E-08 upregulation upregulation TBC domain family member 5 Q80XQ upregulation upregulation pre-mrna processing factor 4B Q E-06 upregulation upregulation Rho guanine nucleotide Q9ES upregulation upregulation exchange factor 7 DNA topoisomerase I Q E-07 upregulation upregulation B cell CLL/lymphoma 9 like Q67FY E-07 upregulation upregulation RAN binding protein 3 Q9CT upregulation upregulation valosin containing protein interacting protein PC4 and SFRS interacting protein exocyst complex component eukaryotic translation elongation factor 2 A-Raf proto-oncogene, serine/threonine kinase Q8CDG E-08 upregulation upregulation Q99JF E-07 upregulation upregulation Q5PPR E-08 upregulation upregulation P E-07 upregulation upregulation P E-08 upregulation upregulation CWC22 spliceosome associated protein homolog Q8C5N E-07 upregulation upregulation membrane associated Q6RHR E-07 upregulation upregulation guanylate kinase, WW and PDZ domain containing HDGF like 2 Q3UMU E-08 upregulation upregulation ataxin 2 like Q7TQH E-07 upregulation upregulation ataxin 2 like Q7TQH E-07 upregulation upregulation CDKN2A interacting protein PWP homolog, endonuclein Q8BI E-08 upregulation upregulation Q99LL upregulation upregulation atrophin O E-08 upregulation upregulation zinc finger (CCCH type), RNA binding motif and rich Q E-07 upregulation upregulation transcription factor 2 Q3UXQ E-09 upregulation upregulation
3 GS(ph)PV TTREPTPR GSGRGGT FT(ph)APR GSPHSEG GKRS(ph)P EPSK GTEKRES( ph)pspapk PR GTSDSSS GNVS(ph)E GDSPPDSQ EDTFHGR GYPSPGA HS(ph)PR HASSS(ph )PESLKPTP APGSR HSSET(ph )FSAPTR HTAPSS(p h)pspgtrt IDASKNE EDEGHSN S(ph)SPR IPRPSVSQ GCS(ph)R ISQHGGS (ph)stslss TK KAGS(ph) DAAASRP R KEVQS(p h)peqvk KEVQS(p h)peqvks EK KLGVSVS (ph)psr KLRSPT(p h)k KNS(ph)F GYR KPIKS(ph) PSKDASSG K KPS(ph)P QPS(ph)PP R KQES(ph) PLLVTK KRS(ph)P SPSPTPEA K nuclear receptor corepressor 2 LSM4A, mrna processing body assembly factor nuclear receptor corepressor 2 repetitive matrix Q9WU E-06 upregulation upregulation Q8K2F upregulation upregulation Q9WU E-06 upregulation upregulation Q52KI E-07 upregulation upregulation PHD finger protein 0 Q9D8M E-07 upregulation upregulation tensin 2 Q8CGB E-08 upregulation upregulation insulin receptor substrate transforming growth factor beta receptor associated protein heterogeneous nuclear ribonucleoprotein D cytoplasmic linker associated protein 2 doublecortin like kinase Q8BTI E-07 upregulation upregulation P E-07 upregulation upregulation Q3UR E-07 upregulation upregulation Q E-05 upregulation upregulation Q8BRT E-07 upregulation upregulation Q9JLM E-09 upregulation upregulation ribosomal protein L6 P E-09 upregulation upregulation BCL2 associated transcription factor BCL2 associated transcription factor zinc finger CCCH-type containing 8 Rho GTPase activating protein 29 La ribonucleoprotein domain family member 4B pre-mrna processing factor 4B Q8K upregulation upregulation Q8K upregulation upregulation Q0P E-07 upregulation upregulation Q8CGF E-07 upregulation upregulation Q6A0A E-08 upregulation upregulation Q E-09 upregulation upregulation cyclin K O upregulation upregulation RUN and cysteine rich domain containing beclin interacting protein SWI/SNF related, matrix associated, actin dependent regulator of chromatin subfamily c member 2 Q80U E-07 upregulation upregulation Q6PDG E-08 upregulation upregulation
4 KSLSDS(p h)esddsk SK KVAPLSS S(ph)LDTS LDFSK KVELS(ph )ESEEDKG SK LASDDRP S(ph)PPR LNHS(ph) PPQSSSR LNRSDS( ph)dsstla K LPLEPET PGSLVGS( ph)pr LRQQSSS SKGDS(ph) PELKPR LRS(ph)PS NDSAHR LRTS(ph) PALK LS(ph)PC LHR LSSLEKSS (ph)ptpr LSSLRAS( ph)tsk LSSLRAS( ph)tskses SQK LSSS(ph)L DNKEK LSSSSS(ph )REPYK LVEPHS(p h)pspssk MSSEGPP RMS(ph)PK NEKS(ph) EEEQSSAS VK NEKS(ph) EEEQSSAS VKK NGHS(ph) PERPTASD CQSPENS HDAGNCS NLMEETK NNPKS(p h)pqkpivr NSQEDSE DS(ph)EEK DVK QEPGGS HMS(ph)E TEDTGR chromobox 3 P E-07 upregulation upregulation UV radiation resistance associated repetitive matrix RNA binding motif protein 0 RNA binding motif protein 26 WW and C2 domain containing 2 SZT2, KICSTOR complex subunit DAB2 interacting protein Q8K E-07 upregulation upregulation A2A8V upregulation upregulation Q99KG upregulation upregulation Q6NZN E-08 upregulation upregulation Q6NXJ E-09 upregulation upregulation A2A9C E-07 upregulation upregulation Q3UHC E-08 upregulation upregulation zinc finger CCCH-type E9Q E-07 upregulation upregulation containing 3 centrosomal protein 70 Q6A E-07 upregulation upregulation family with sequence similarity 7 member A Q7TNF E-09 upregulation upregulation eukaryotic translation P E-07 upregulation upregulation elongation factor delta ribosomal protein S6 P E-07 upregulation upregulation ribosomal protein S6 P E-06 upregulation upregulation protein phosphatase regulatory subunit 2A Q9DBR E-08 upregulation upregulation YTH domain containing E9Q5K E-0 upregulation upregulation zinc finger protein 609 Q8BZ E-08 upregulation upregulation ataxin 2 O E-07 upregulation upregulation heterogeneous nuclear ribonucleoprotein C (C/C2) Q9Z upregulation upregulation heterogeneous nuclear Q9Z E-07 upregulation upregulation ribonucleoprotein C (C/C2) golgin A2 A2AN E-06 upregulation upregulation B-Raf proto-oncogene, serine/threonine kinase nuclear casein kinase and cyclin dependent kinase substrate G-patch domain containing 8 P upregulation upregulation Q80XU E-05 upregulation upregulation A2A6A E-08 upregulation upregulation
5 QEQLS(ph )PR QGSGRES (ph)pslvs R QKIDDRD S(ph)EEEG PSNQR QQASKS( ph)pyngv R QSEES(ph )PADDGEL RR QSVDKV TS(ph)PTK V QVQQNS( ph)lhr RAKS(ph) PTPDGSER RAS(ph)D GGANIQL HAQQLLK RAS(ph)S EIVTEGK RAVVVS( ph)pkeen K RDEQAA S(ph)DPM DQETVSR RDS(ph)F SENEK RDSSES(p h)qlaste SDKPTTGR RGFS(ph) DSGGGPP AK RHS(ph)S PSSPTSPK RHSS(ph) PSSPTSPK RIQQQLG EEAS(ph)P R RKPS(ph) PEPEGEVG PPK RLPSSPAS (ph)pspk RLS(ph)P VEK RNS(ph)S SSSSPSERP R RPAS(ph) VSSAAAE HEAR nuclear receptor F8VQL E-06 upregulation upregulation corepressor 2 ataxin 2 like Q7TQH upregulation upregulation DEAD-box helicase 54 Q8K4L E-08 upregulation upregulation cyclin L Q52KE E-07 upregulation upregulation ATRX, chromatin remodeler Q E-07 upregulation upregulation caldesmon D3Z6I E-07 upregulation upregulation intersectin Q9Z0R E-07 upregulation upregulation YTH domain containing E9Q5K E-07 upregulation upregulation SIK family kinase 3 Q6P4S upregulation upregulation SUMO/sentrin specific Q8BUH E-08 upregulation upregulation peptidase 7 cyclin K O E-08 upregulation upregulation MON2 homolog, regulator of endosometo-golgi trafficking CWC22 spliceosome associated protein homolog chromosome 8 open reading frame 25 pinin, desmosome associated protein cyclin dependent kinase 4 cyclin dependent kinase 4 Rho guanine nucleotide exchange factor 40 Q80TL E-07 upregulation upregulation Q8C5N E-0 upregulation upregulation Q8BH upregulation upregulation O upregulation upregulation O E-08 upregulation upregulation O E-08 upregulation upregulation Q3UPH E-06 upregulation upregulation interferon regulatory factor 2 binding protein 2 E9QP upregulation upregulation erythrocyte membrane A2AUK E-08 upregulation upregulation protein band 4. like ETS variant 6 P E-09 upregulation upregulation echinoderm microtubule associated protein like 3 interferon regulatory factor 2 binding protein 2 Q8VC E-08 upregulation upregulation E9QP E-06 upregulation upregulation
6 RPGS(ph) VSSTDQER RPMEED GEEKS(ph) PSK RPMEED GEEKS(ph) PSK RQDTTRS (ph)pslap TQR RQMS(ph) LTEK RSEAEEG EVRT(ph)P TK RVSSET(p h)hqgpgt PESK RYS(ph)G DSDSSASS AQSGPMG AR S(ph)NEQ MVSEKPSE SK S(ph)PVK QDKSEIST DPK S(ph)QPC VLNDKK S(ph)VSTL RPCAK SAS(ph)E PSLHR SAVRPSP S(ph)PER SCS(ph)FS SESR SES(ph)LS NCSIGKK SGGPNSC KSDDYMP MS(ph)PTS VSAPK SGSS(ph)P EMKDKPR SGSSQEL DGKPSAS( ph)pqer SHLETM GSS(ph)PL STTK SKPNVPA ES(ph)R SKS(ph)E DMDSVES K SKS(ph)Q SSGSSATH interferon regulatory factor 2 binding protein like interleukin enhancer binding factor 3 interleukin enhancer binding factor 3 Q8K3X E-08 upregulation upregulation Q9ZX E-06 upregulation upregulation Q9ZX E-08 upregulation upregulation tensin 2 Q8CGB E-08 upregulation upregulation DAB2 interacting protein erythrocyte membrane protein band 4. like nuclear mitotic apparatus protein growth arrest specific 2 like Q3UHC E-06 upregulation upregulation A2AUK E-07 upregulation upregulation E9Q7G E-05 upregulation upregulation Q8JZP E-09 upregulation upregulation calpastatin P E-08 upregulation upregulation family with sequence similarity 53 member B MON2 homolog, regulator of endosometo-golgi trafficking A-Raf proto-oncogene, serine/threonine kinase DENN domain containing 4C ankyrin repeat and sterile alpha motif domain containing A insulin receptor substrate 2 serine/threonine kinase interacting protein serine and arginine rich splicing factor 4 activating transcription factor 7 interacting protein insulin receptor substrate 2 Q8BTI upregulation upregulation Q8BGR E-08 upregulation upregulation Q80TL E-08 upregulation upregulation P upregulation upregulation Q8BTI E-07 upregulation upregulation A6H8H upregulation upregulation P upregulation upregulation P E-06 upregulation upregulation Q8BTI E-07 upregulation upregulation Q8BTI E-08 upregulation upregulation Q3TAA E-07 upregulation upregulation Q8VE E-6 upregulation upregulation Q7TT E- upregulation upregulation P upregulation upregulation
7 PISVPGAR SPERTEE VLSPDGSP SKSPS(ph) K SPQESTG DPGNSSSV SDGKGSSE RGS(ph)PIE K SRS(ph)PF ASTR SRTGS(ph )ESSQTGA SATSGR SRTPS(ph) ASHEEQQ E SS(ph)FSN SADDIKSK SS(ph)SK DSRPSQA AGDNQG DEAK SS(ph)SSA GSEVGGQ STGSNHK SSESVVD EDGGRS(p h)pr SSS(ph)LP SDRGPPA R STS(ph)L NERPK SVGKVEP SSQSPGRS( ph)pr T(ph)AEE EDEADPK R T(ph)ASN AEQYKYG K T(ph)KSD ESGEEKN GDEDCQR TAS(ph)E GSEAETPE APKQPAK K TAVSVA QGGHS(ph )R TDGS(ph)I S(ph)GDR QPVTVAD YISR TDSEKPF RGSQS(ph) PK adducin 3 Q9QYB E-05 upregulation upregulation transcriptional repressor GATA binding praja ring finger ubiquitin ligase eukaryotic translation initiation factor 4B Q925H E-07 upregulation upregulation O E-08 upregulation upregulation Q8BGD E-08 upregulation upregulation small glutamine rich tetratricopeptide repeat containing alpha Q8BJU upregulation upregulation chromobox 5 Q E-06 upregulation upregulation thyroid hormone receptor associated protein 3 trinucleotide repeat containing 6B microtubule associated serine/threonine kinase 3 DENN domain containing 4C Q569Z E-06 upregulation upregulation Q8BKI E-07 upregulation upregulation Q3U E-08 upregulation upregulation A6H8H upregulation upregulation TSC complex subunit 2 Q E-06 upregulation upregulation ELKS/RAB6- interacting/cast family member Q99MI upregulation upregulation parathymosin Q9D0J E-07 upregulation upregulation actin filament associated protein La ribonucleoprotein domain family member La ribonucleoprotein domain family member 7 Q80YS upregulation upregulation Q6ZQ E-07 upregulation upregulation Q05CL E-05 upregulation upregulation PBX homeobox 2 O E-09 upregulation upregulation FMR autosomal homolog 2 thyroid hormone receptor associated protein 3 Q6P5B downregulation upregulation Q569Z upregulation upregulation
8 TEEVLSP DGSPSKS( ph)pskk TFS(ph)A TVR TGS(ph)N ISGASSDV SLDEQYK HQLEETK TGS(ph)T GALGPSER TKEYVSN DATQS(ph) DDEEKLQ SQQTDTD GGR TS(ph)SD PNLNNHS QEVR TSS(ph)D PNLNNHS QEVR TSS(ph)T NEDEDLN PEQKIER TTS(ph)PT AESVRPTE SMRR TVIRLPSG SGPAS(ph) PTTGSAV DIR VALGDG VQLPPGD Y(ph)STTP GGTLFSTT (ph)pggtr VCS(ph)P YNHR VEPSSQS PGRS(ph)P R VEQATKP S(ph)FESG R VKTVIS(p h)pr VLAPCS( ph)pseerr VLS(ph)P PHTK VPS(ph)P DHRR VRAHS(p h)paegas SESSSPGP K VRHDT(p h)pdps(ph) PPR adducin 3 Q9QYB upregulation upregulation calcium regulated heat stable protein oxysterol binding protein protein phosphatase regulatory subunit 2C Q9CR upregulation upregulation Q3B7Z E-07 upregulation upregulation Q3UMT E-06 upregulation upregulation WD repeat domain 44 Q6NVE upregulation upregulation myotubularin related protein 4 myotubularin related protein 4 Q9XS E-07 upregulation upregulation Q9XS E-07 upregulation upregulation transcription factor 2 Q3UXQ E-07 upregulation upregulation lipase E, hormone sensitive type P upregulation upregulation AHNAK nucleoprotein E9Q upregulation upregulation eukaryotic translation initiation factor 4E binding protein TOP binding arginine/serine rich protein ELKS/RAB6- interacting/cast family member RNA binding motif protein, X-linked like- inositol-trisphosphate 3- kinase B Q upregulation upregulation Q80Z E-08 upregulation upregulation Q99MI E-08 upregulation upregulation Q9VM E-06 upregulation upregulation Q8BTI upregulation upregulation B2RXC E-08 upregulation upregulation ubiquitin associated protein Q8BH E-09 upregulation upregulation pre-mrna processing Q4FK E-08 upregulation upregulation factor 38A folliculin Q8QZS E-07 upregulation upregulation BUD3 homolog Q8R E-08 upregulation upregulation
9 VSS(ph)ET HQGPGTP ESK VTRS(ph) QEELREEK VVSS(ph) TSEEEEAF TEK nuclear mitotic apparatus protein E9Q7G E-0 upregulation upregulation afadin, adherens junction formation factor Q9QZQ E-05 upregulation upregulation oxidation resistance E9Q0A E-06 upregulation upregulation Phosphopeptides regulated by both SSE and insulin. 3T3-L adipocytes at the 4th day of differentiation were treated with SSE (70 µg/ml) or insulin (00 nm) for 30 min. Phosphoproteomic analysis was performed as described in Materials and Methods. Only phosphopeptides that were significantly affected by both SSE and insulin are presented. P-value was calculated by Student s t- test, compared to control, untreated cells. Phospho Site AGETRFT(ph)DTR AQKPPS(ph)PPAMEN GTR DDSHSAEDS(ph)EDE KDDHK EEALDEAEEPES(ph)P PPPPRS(ph)PS(ph)PEPT VVDTPSHASQSAR EENVNS(ph)PEDKR EEQSGPVDEKGNDS(p h)dgeaesddpekk EGKEDEAS(ph)TDVD EKPK ELSSEPRT(ph)PPAQK ERGT(ph)PPVDPK ESEALPEKEGDELGEG ERPEDDTAAIELS(ph)S( ph)deaveveevieesr EYGS(ph)TSSIDR FHDS(ph)EGDDT(ph)E ETEDYR Table S2. Phosphopeptides regulated by SSE, but not affected by insulin. Entrez Gene Name ID Phospho Fold Change eukaryotic translation elongation factor 2 trna methyltransferase nuclear casein kinase and cyclin dependent kinase substrate arginine-glutamic acid dipeptide repeats ATRX, chromatin remodeler vacuolar protein sorting 4 homolog B solute carrier family 6 member microtubule associated protein A proline rich coiled-coil 2A caveolae associated protein signal induced proliferation associated like 2 BCL2 associated transcription factor p-value Regulation by SSE P E-05 upregulation Q3TX E-06 upregulation Q80XU E-07 upregulation Q80TZ E-06 upregulation Q E-06 upregulation P E-09 upregulation P E-09 upregulation Q9QYR upregulation Q7TSC E-07 upregulation O downregulation Q80TE E-06 upregulation Q8K E-05 upregulation FKAEAPLPS(ph)PK AHNAK nucleoprotein E9Q upregulation FLMECRNS(ph)PVAK GAAAADLLSS(ph)SPE SQHGGTQPPGGGQPL LQPTK GDS(ph)PELKPR GEEGS(ph)EEEETENG PKPK GGGSGGGDES(ph)EG EEVDED GHYEVTGS(ph)DDEA GK eukaryotic translation initiation factor 4E binding protein Q upregulation protein tyrosine Q6PB E-06 upregulation phosphatase, nonreceptor type 23 DAB2 interacting Q3UHC E-05 upregulation protein CTR9 homolog, Q E-07 upregulation Paf/RNA polymerase II complex component purine rich element O upregulation binding protein B AHNAK nucleoprotein E9Q E-06 upregulation
10 GLS(ph)VDSAQEVKR HADGEKEDQFNGS(p h)pprpqpr HSLDS(ph)DEEDDDEE GSSK IEDVGS(ph)DEEDDS(p h)gkdkk transformation/transcri ption domain associated protein vir like m6a methyltransferase associated CD2 cytoplasmic tail binding protein 2 heat shock protein 90 alpha family class B member Q80YV E-06 upregulation A2AIV upregulation Q9CWK E-08 upregulation P E-05 upregulation ILEQQNSSRT(ph)LEK septin 7 O upregulation KEES(ph)EES(ph)DDD MGFGLFD KPAATS(ph)PLSPMA NGGR KSRVS(ph)VS(ph)PGR KSS(ph)SSSGVPYSPAI PNK ribosomal protein lateral stalk subunit P2 pleckstrin homology like domain family B member repetitive matrix exocyst complex component 7 P downregulation Q6PDH E-07 upregulation A2A8V upregulation O E-06 upregulation KTS(ph)GPPVSELITK histone cluster, He P E-08 upregulation LNHVAAGLVS(ph)PS LK LRMS(ph)GAGK LVLS(ph)GEKK MLAESDDS(ph)GDEE SVSQTDKTELQSTLR QASTDAGT(ph)AGAL TPQHVR serine and arginine rich splicing factor phosphofructokinase, liver type eukaryotic translation initiation factor 4 gamma 3 oxysterol binding protein Yes associated protein E9Q6E upregulation P E-05 upregulation Q80XI upregulation Q3B7Z E-06 upregulation P upregulation QGS(ph)PTPALPEKR tensin A0A087W upregulation Q94 QGSGRESPS(ph)LVSR ataxin 2 like Q7TQH downregulation QQSHFAMMHGGTGF AGIDSSS(ph)PEVK RADANLLTDTGTESS PRSPVCS(ph)LR RALS(ph)PLPAR RKAS(ph)GPPVSELITK RKS(ph)ELGAEPGHFG VCVDSLTSDK RLS(ph)SLRAST(ph)SK RPPS(ph)PDPNTKVSE EAESQQWDTSK RRDS(ph)YYDR RS(ph)PERPTGDLR RTAS(ph)AGTVSDAE AR RVES(ph)EESGDEEGK K RVESEES(ph)GDEEGK K poly(rc) binding P upregulation protein exosome component 5 Q9CRA E-05 upregulation pleckstrin homology like domain family B member Q6PDH upregulation histone cluster H P E-08 upregulation family member c myosin IXB Q9QY E-07 upregulation ribosomal protein S6 P upregulation spectrin beta, nonerythrocytic transformer 2 alpha homolog zinc finger CCCH-type containing 3 Ras association (RalGDS/AF-6) and pleckstrin homology domains protein phosphatase regulatory subunit 7 protein phosphatase regulatory subunit 7 Q E-05 upregulation A0A0N4S E-06 upregulation VC2 E9Q E-06 upregulation F2Z3U E-07 upregulation Q3UM E-06 upregulation Q3UM E-07 upregulation
11 RVQT(ph)PLLR septin 9 Q80UG E-07 upregulation S(ph)LPTSPERR protein phosphatase regulatory subunit 3D SANDSTVHS(ph)PFTK SANQSPQSVGGS(ph) GIDS(ph)GVESTSDSLR SDDYMPMS(ph)PTSVS APK SDDYMPMS(ph)PTSVS APK SLPASGTPQS(ph)PPA VK A2AJW E-06 upregulation ubiquitin-associated Q80X E-06 upregulation protein 2-like lipin Q9ZP E-0 upregulation insulin receptor substrate 2 insulin receptor substrate 2 Ras association (RalGDS/AF-6) and pleckstrin homology domains SLPTS(ph)PERR protein phosphatase regulatory subunit 3D SPS(ph)ASSLSSMSSVA SSVSSKPSR SQDATVSPGSEQSEKS (ph)pgpivsr SQSRS(ph)HS(ph)PLP APPSK SRTESITATS(ph)PASM VGGKPGSFR SSS(ph)PEDRYTEQER STPSQVTSEEKDGHS( ph)pmsk STSAPQMSPGS(ph)S(p h)dnqssspqpaqqk SVGKVEPSSQS(ph)PG R SVTPDSLGHT(ph)PPA R TAS(ph)EGDGGAAGG AGTAGGRPMSVAGSP LSPGPVR TAS(ph)FSESRADEVA PAK TAS(ph)FSESRADEVA PAKK TDGFAEAIHS(ph)PQV AGVPR TEEVLSPDGSPSKS(ph) PSK TEHAPS(ph)PSSGGTV K TELS(ph)PRPGAAGR TPLGAS(ph)LDEQSSG TPK CAP-Gly domain containing linker protein zinc finger C3HC-type containing serine and arginine rich splicing factor 6 insulin receptor substrate CWC22 spliceosome associated protein homolog microtubule associated protein A ubiquitin-associated protein 2-like ELKS/RAB6- interacting/cast family member insulin receptor substrate insulin receptor substrate 2 P E-08 upregulation P upregulation F2Z3U downregulation A2AJW upregulation Q922J downregulation Q80YV upregulation Q3TWW E-06 upregulation P upregulation Q8C5N E-08 upregulation Q9QYR E-06 upregulation Q80X E-09 upregulation Q99MI E-07 upregulation P E-05 upregulation P E-05 upregulation ATP citrate lyase Q3TS upregulation ATP citrate lyase Q3TS upregulation translocated promoter region, nuclear basket protein F6ZDS upregulation adducin 3 Q9QYB upregulation cyclin dependent kinase 3 eukaryotic translation initiation factor 2B subunit delta sperm specific antigen 2 Q69ZA upregulation Q upregulation Q922B upregulation VGGS(ph)SVDLHR catenin delta P E-05 upregulation VPELDGAGSTEQDKS HS(ph)NSSTLSDR VTNDSSSSSSSSSSDSD S(ph)DGEEHDSDRAPR Ral GTPase activating protein catalytic alpha subunit 2 NADH:ubiquinone oxidoreductase subunit V3 A3KGS E-07 upregulation Q3U E-08 upregulation
12 YCRPESQEHPEADPG SAAPY(ph)LK signal transducer and activator of transcription 3 P upregulation Phosphopeptides regulated by SSE, but not affected by insulin. 3T3-L adipocytes at the 4th day of differentiation were treated with SSE (70 µg/ml) or insulin (00 nm) for 30 min. Phosphoproteomic analysis was performed as described in Materials and Methods. Only phosphopeptides that were significantly affected by SSE, but not affected by insulin are presented. P-value was calculated by Student s t-test, compared to control, untreated cells. Phospho Site AASDGQYENQS(ph)PE ATSPR AAS(ph)DGQYENQSPE ATSPR Table S3. Phosphopeptides regulated by insulin, but not affected by SSE. Entrez Gene Name ID Phospho Fold Change tensin A0A087W Q94 tensin A0A087W Q94 HAAYGGYST(ph)PEDR tensin A0A087W Q94 RAAS(ph)DGQYENQSP EATSPR EAFEEMEGTSPSS(ph)PP HSVAR HPGAHQGNLASSLHSN AVISPGS(ph)PSLGR IEQNLQS(ph)PTQQQTA R LPS(ph)SEIHPEESLYR VQEESYPQS(ph)PR APS(ph)LTSDSEGKKPA QAVK SPDPEMVPRGS(ph)PVR tensin tensin tensin unc-5 like autophagy activating kinase TBC domain family member 6 BCLAF and THRAP3 family member 3 SEC6 homolog A, endoplasmic reticulum export factor SEC6 homolog A, endoplasmic reticulum export factor YSEPERPSS(ph)R SEC6 homolog A, endoplasmic reticulum export factor RKPGAGGS(ph)PALAR MAP7 domain containing TGS(ph)TSSKEDDYESD AATIVQK TS(ph)PADHGGSVGSES GGSAVDSVAGEHSVSGR NGHSPERPTAS(ph)DC QSPENSHDAGNCSNLM EETK HMSES(ph)PNRKVEK KADREQS(ph)PVSK QGDETPSTNNGS(ph)D DEKTGLK IHRAS(ph)DPGLPAEEP KEKPPR A0A087W Q94 A0A087W Q94 A0A087W Q94 A0A0R4J0 B3 p-value Regulation by Insulin upregulation upregulation upregulation upregulation upregulation upregulation upregulation A2ABG upregulation A2AG upregulation A2AIX upregulation A2AIX upregulation A2AIX upregulation A2AJI upregulation ubiquitin protein ligase E3 component n-recognin 4 A2AN upregulation ubiquitin protein A2AN upregulation ligase E3 component n-recognin 4 golgin A2 A2AN upregulation peptidylprolyl isomerase G peptidylprolyl isomerase G RAB GTPase activating protein carbamoylphosphate synthetase A2AR upregulation A2AR upregulation A2AWA upregulation B2RQC upregulation
13 IHRAS(ph)DPGLPAEEP K ARSPS(ph)PCPFR VTSSMSS(ph)PSMQPK GAPHTVS(ph)HEDIR CETS(ph)PPSS(ph)PRPL RLDR DMS(ph)PTSTDTEVHR RETVVESQSSQSPS(ph)P KR DQKEEGNDYDT(ph)R AEAPLPS(ph)PK ISMPDIDLHLKS(ph)PK GDLGASS(ph)PSMK LRS(ph)EDGVEGDLGET QSR SKGHYEVTGS(ph)DDE AGKLQGSGVSLASK VNGDDHHEEDMDMS( ph)d EEKRPTEAVS(ph)PK DYDEEEQGYDS(ph)EKE K VNGDDHHEEDMDMS( ph)d VS(ph)SETHQGPGTPES K VSSETHQGPGT(ph)PES K VSS(ph)ETHQGPGTPES KK 2, aspartate transcarbamylase, and dihydroorotase carbamoylphosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase inositoltrisphosphate 3- kinase B PTPRF interacting protein alpha PTPRF interacting protein alpha PTPRF interacting protein alpha AHNAK nucleoprotein 2 SR-related CTD associated factor YTH domain containing AHNAK nucleoprotein AHNAK nucleoprotein AHNAK nucleoprotein AHNAK nucleoprotein AHNAK nucleoprotein serine and arginine rich splicing factor serine and arginine rich splicing factor serine and arginine rich splicing factor serine and arginine rich splicing factor nuclear mitotic apparatus protein nuclear mitotic apparatus protein nuclear mitotic apparatus protein B2RQC upregulation B2RXC upregulation B2RXQ upregulation B2RXQ upregulation B2RXQ upregulation E9PYB upregulation E9PZM upregulation E9Q5K upregulation E9Q upregulation E9Q upregulation E9Q upregulation E9Q upregulation E9Q upregulation E9Q6E upregulation E9Q6E upregulation E9Q6E upregulation E9Q6E upregulation E9Q7G upregulation E9Q7G upregulation E9Q7G upregulation LKQTENAFS(ph)PSR caldesmon E9QA upregulation DTTQRS(ph)PVHVQPT AR GSPAPSSASSSASDLS(ph )RSPAHSR YS(ph)PPRDDDKVDNQ AK TLS(ph)SSRPPLLLR GRAS(ph)PGGVSTSSSD GKAEK SSSAGKES(ph)PKVR RAQST(ph)DSLGTSSSL QSK AHNAK nucleoprotein 2 F7DBB upregulation G protein pathway G3UXW upregulation suppressor thyroid hormone G5E upregulation receptor interactor 2 eukaryotic O upregulation elongation factor 2 kinase atrophin O upregulation cyclin dependent kinase 4 rabaptin, RAB GTPase binding effector protein O upregulation O upregulation
14 GFS(ph)DSGGGPPAK pinin, desmosome O upregulation associated protein SRPLS(ph)MDAR serine/threonine O upregulation kinase 0 SPRPGSS(ph)VPEHSPR unc-5 like O upregulation autophagy activating kinase RMS(ph)LEGR caspase 8 O upregulation YSPTS(ph)PK RNA polymerase II subunit A EIS(ph)DDEAEEEKGEK heat shock protein 90 alpha family class B member IEDVGS(ph)DEEDDSGK DKK KVS(ph)ADGAAKAEPK SMS(ph)ADEDLQEPSR heat shock protein 90 alpha family class B member high mobility group nucleosome binding domain negative elongation factor complex member E P upregulation P upregulation P upregulation P upregulation P upregulation RKS(ph)LSDSESDDSK chromobox 3 P upregulation SKVGS(ph)TENIK VGS(ph)TENIKHQPGG GR SKVGS(ph)TENIKHQPG GGR SLS(ph)QRSR EEQTDTS(ph)DGESVTH HIR QIS(ph)EDVDGPDNR microtubule associated protein 4 microtubule associated protein 4 microtubule associated protein 4 natural killer cell triggering receptor G protein nucleolar (putative) neural precursor cell expressed, developmentally down-regulated 4 P upregulation P upregulation P upregulation P upregulation P upregulation P upregulation AGS(ph)DAAASRPR ribosomal protein L6 P upregulation KEES(ph)EESEDDMGFG LFD NKST(ph)ESLQANVQR GRASS(ph)HSSQSQGGG SVTK ribosomal protein, large, P P upregulation ribosomal protein P upregulation L3 lamin A/C P upregulation LDNARQS(ph)AER lamin A/C P upregulation SES(ph)PKEPEQLRK YVDSEGHLYT(ph)VPIR STYQDKPST(ph)PAEKK KQQQEPTCEPS(ph)PK heterogeneous nuclear ribonucleoprotein A P upregulation caveolin P upregulation calpastatin P upregulation high mobility group AT-hook 2 P upregulation SRSLS(ph)ASPALGSTK NAD kinase P upregulation VSGAGLS(ph)PSRK DVGHLEEGASGGLLS(p h)pstphsr S(ph)PVGDTGLGKR tankyrase binding protein tankyrase binding protein tankyrase binding protein P upregulation P upregulation P upregulation S(ph)HTGEAAAVR BCL2 like 3 P upregulation
15 LKAGGS(ph)VESLR SAM and SH3 domain containing S(ph)LTEGEMKK SAM and SH3 domain containing VISS(ph)IEQK tyrosine 3- monooxygenase/tryp tophan 5- monooxygenase activation protein gamma VISSIEQKTS(ph)ADGNE K LSS(ph)LRAS(ph)TSKSE SSQK P upregulation P upregulation P upregulation tyrosine 3- monooxygenase/tryp tophan 5- monooxygenase activation protein gamma P upregulation ribosomal protein S6 P upregulation LSS(ph)LRASTSK ribosomal protein S6 P upregulation NYQQNYQNSESGEKNE GSES(ph)APEGQAQQR NYQQNYQNSES(ph)GE KNEGSESAPEGQAQQR RSPS(ph)PYYSR TAS(ph)EGDGGAAGGA GTAGGRPMSVAGSPLS(p h)pgpvr TAS(ph)EGDGGAAGGA GTAGGRPMSVAGSPLS(p h)pgpvr TAS(ph)EGDGGAAGGA GTAGGRPMSVAGS(ph)P LSPGPVR KADS(ph)DSEDKGEESK PK LMELHGEGGSS(ph)GK FLTQS(ph)PK STANHDSES(ph)PVHSP SAHR MDIDRS(ph)PGLLGTPD LK TTSRS(ph)PVLSR Y-box binding protein Y-box binding protein transformer 2 beta homolog insulin receptor substrate 2 insulin receptor substrate 2 insulin receptor substrate 2 P upregulation P upregulation P upregulation P upregulation P upregulation P upregulation chromobox P upregulation ribosomal protein S3A lysosomal trafficking regulator lysosomal trafficking regulator myosin phosphatase Rho interacting protein mitogen-activated protein kinase kinase kinase kinase 4 GSSNKDFT(ph)PGRDK RB binding protein 6, ubiquitin ligase SRS(ph)PQAFR RB binding protein 6, ubiquitin ligase LERT(ph)PEKDK RB binding protein 6, ubiquitin ligase IHVS(ph)DQELQSANAS VDDSRLEELK SRNS(ph)PLLDR TAS(ph)EGS(ph)EAETPE APKQPAKK ubiquitin like modifier activating enzyme microtubule affinity regulating kinase 2 La ribonucleoprotein domain family member 7 P upregulation P upregulation P upregulation P upregulation P upregulation P upregulation P upregulation P upregulation Q upregulation Q upregulation Q05CL upregulation
16 TAS(ph)EGSEAETPEAP KQPAK TARPNSEAPLS(ph)GSE DADDSNKLSKK TARPNSEAPLS(ph)GSE DADDSNKLSK MGS(ph)PKPER VQSQEETRS(ph)DEEDR ASEPK KKLGVSVS(ph)PSR GGS(ph)REYETGGGSSS SR RGS(ph)PVPPVPERR KGSPT(ph)PAYPER EKEPEAASSRGS(ph)PV R VGDTEKPEPERS(ph)PP NR KAKPAMPQDSVPS(ph) PR KSYESSEDCPEAASS(ph) PTRK GS(ph)PQQIDHAK HGSGPNIILTGDSS(ph)P GFSK SPS(ph)APVCK KSRS(ph)DNALNLVTER SDCQRS(ph)PPGVLK STS(ph)VDDTDKSSSEAI MVR KREQT(ph)ASAPATPLV SK QDS(ph)NPKPKPSNEIT R LPEKEPACTYGNNVPLS (ph)pvdgsnknpaasyl K TRLAS(ph)ESANDDNE DS La ribonucleoprotein domain family member 7 Q05CL upregulation eukaryotic Q05D upregulation translation initiation factor 5B eukaryotic Q05D upregulation translation initiation factor 5B zinc finger CCCHtype Q0P upregulation containing 8 zinc finger CCCHtype Q0P upregulation containing 8 zinc finger CCCHtype Q0P upregulation containing 8 RNA binding motif Q0VBL upregulation protein 5 family with sequence Q48V upregulation similarity 83 member H family with sequence Q48V upregulation similarity 83 member H ubiquitin specific Q3TIX upregulation peptidase 39 chromosome 9 open Q3TQI upregulation reading frame 78 ATP citrate lyase Q3TS upregulation alkb homolog 5, RNA demethylase KH-type splicing regulatory protein CREB regulated transcription coactivator 2 family with sequence similarity 7 member B prickle planar cell polarity protein TRAF-type zinc finger domain containing cordon-bleu WH2 repeat protein like cordon-bleu WH2 repeat protein like cordon-bleu WH2 repeat protein like cordon-bleu WH2 repeat protein like Q3TSG upregulation Q3U0V upregulation Q3U upregulation Q3U3E upregulation Q3U5C upregulation Q3UDK upregulation Q3UMF upregulation Q3UMF upregulation Q3UMF upregulation Q3UMF upregulation HDGF like 2 Q3UMU upregulation S(ph)EGLSLER HDGF like 2 Q3UMU upregulation AQEDGQDS(ph)EDGPR HDGF like 2 Q3UMU upregulation GGSSEELHDSPR LAS(ph)ESANDDNEDS HDGF like 2 Q3UMU upregulation HTAPSS(ph)PSPGTR transforming growth Q3UR upregulation factor beta receptor associated protein MSGS(ph)PGVSR splicing factor SWAP Q3USH upregulation
17 S(ph)NSLPHSAVSNAAS K VPS(ph)PDHR QS(ph)PSPSTRPIR TRHS(ph)PTPQQSNR SS(ph)SKDSRPSQAAGD NQGDEAKEQTFSGGTSQ DIK ERS(ph)PALK NKKS(ph)PEIHR RSSMSSCGSSGYFSSSPT( ph)ls(ph)ssppvlcnpk GPRPCTS(ph)PQPLR SDDSKSSS(ph)PEPVTHL K VHAS(ph)RPASLDSGR ESDTKNEVNGTSEDIKS EGDTQS(ph)N LNS(ph)DEEGESSGKR LNS(ph)DEEGESSGK KLNS(ph)DEEGESSGKR SAEPPRS(ph)PLLKR RES(ph)PPIR GGT(ph)YPRR TLS(ph)PGRR IGS(ph)DPLAYEPK YSDEQLNSGRQS(ph)PS QNER HAEQNGPVDGQGDNP GSQAAEHGADT(ph)AV PSDGDK WD repeat domain 20 pre-mrna processing factor 38A repetitive matrix repetitive matrix thyroid hormone receptor associated protein 3 thyroid hormone receptor associated protein 3 thyroid hormone receptor associated protein 3 DEP domain containing MTOR interacting protein PEAK related, kinase-activating pseudokinase Rho GTPase activating protein coiled-coil domain containing 88A LUC7 like 3 premrna splicing factor LEO homolog, Paf/RNA polymerase II complex component LEO homolog, Paf/RNA polymerase II complex component LEO homolog, Paf/RNA polymerase II complex component microtubule associated serine/threonine kinase 2 nuclear receptor corepressor mitogen-activated protein kinase kinase kinase 3 pre-mrna processing factor 4B solute carrier family 9 member A APC, WNT signaling pathway regulator heat shock protein family A (Hsp70) member 4 Q3UWE upregulation Q4FK upregulation Q52KI upregulation Q52KI upregulation Q569Z upregulation Q569Z upregulation Q569Z upregulation Q570Y upregulation Q57I upregulation Q5FWK upregulation Q5SNZ upregulation Q5SUF upregulation Q5XJE upregulation Q5XJE upregulation Q5XJE upregulation Q upregulation Q upregulation Q upregulation Q upregulation Q upregulation Q upregulation Q upregulation SS(ph)FSNSADDIK chromobox 5 Q upregulation KSS(ph)FSNSADDIKSK chromobox 5 Q upregulation SEMKTELS(ph)PRPGAA GR eukaryotic translation initiation Q upregulation
18 factor 2B subunit delta DLKPST(ph)PR nucleophosmin Q upregulation LHQLAMQQSHFPMTH GNTGFSGIESSS(ph)PEVK LHQLAMQQSHFPMTH GNTGFSGIESSS(ph)PEVK TS(ph)PDTLR TS(ph)PDTLRR poly(rc) binding protein 2 poly(rc) binding protein 2 serine and arginine rich splicing factor 2 serine and arginine rich splicing factor 2 Q upregulation Q upregulation Q upregulation Q upregulation LRS(ph)PFLQK drebrin like Q upregulation QLT(ph)QPETSYGR drebrin like Q upregulation SVS(ph)VDSGEQR HTSS(ph)PEVVAEDR LGS(ph)LSAR ADTS(ph)PTYVR KLS(ph)VGGDSDPPLKR B cell CLL/lymphoma 9 like RHO family interacting cell polarization regulator centrosomal protein 70 zinc finger protein 29 zinc finger CCCHtype containing A Q67FY upregulation Q68FE upregulation Q6A upregulation Q6IQX upregulation Q6NZF upregulation RSS(ph)PLLR histone deacetylase 4 Q6NZM upregulation RLNHS(ph)PPQSSSR EAEQGS(ph)GEEKEEKE GDLK AAIS(ph)QLR EQGVESPGAQPASS(ph) PR ERSMS(ph)ENAVR TLTDEVNS(ph)PDSDRR EVAES(ph)PRPR ESTESSNTT(ph)IEDEDV KAR ESTESSNTT(ph)IEDEDV K KDS(ph)QNSSQHSVSSH R KDSQNSS(ph)QHSVSSH R RNA binding motif protein 26 Q6NZN upregulation ATP binding cassette Q6P downregulation subfamily F member oxidative stress Q6P9R upregulation responsive taxilin alpha Q6PAM upregulation mitochondrial fission factor SWI/SNF related, matrix associated, actin dependent regulator of chromatin subfamily c member 2 pleckstrin homology like domain family B member calcium/calmodulin dependent protein kinase II delta calcium/calmodulin dependent protein kinase II delta membrane associated guanylate kinase, WW and PDZ domain containing membrane associated guanylate kinase, WW and PDZ domain containing Q6PCP upregulation Q6PDG upregulation Q6PDH upregulation Q6PHZ upregulation Q6PHZ upregulation Q6RHR upregulation Q6RHR upregulation
19 GVAPADS(ph)PDAPRR TGTGSPFAGNS(ph)PAR S(ph)LPTTVPES(ph)PNY R S(ph)LPTTVPESPNY(ph) R SSPSTGS(ph)LDSGNESK EK AEVSTIHLQS(ph)PGRK SRS(ph)LGGAVGSASGG R ERSDS(ph)GGSSSEPFER SDS(ph)GGSSSEPFER AAAS(ph)PLGESGPSIH PHDK S(ph)KSEDMDSVESK RIS(ph)QSQPVR VAS(ph)DTEDTDRITSK EKVAS(ph)DTEDTDRIT SK GKAS(ph)PFEEDQNR SHS(ph)DTNIASR RSSFS(ph)EGQTAPVAS GTKK RSS(ph)FSEGQTAPVAS GTKK SRS(ph)PIER SANDSTVHS(ph)PFTKR ankyrin repeat and IBR domain containing Q6ZPS upregulation zinc finger CCCHtype Q6ZPZ upregulation containing 4 La ribonucleoprotein Q6ZQ upregulation domain family member La ribonucleoprotein Q6ZQ upregulation domain family member ArfGAP with coiledcoil, Q6ZQK upregulation ankyrin repeat and PH domains 2 synemin Q70IV upregulation zinc and ring finger 2 Q7FD upregulation proline rich coiledcoil 2A proline rich coiledcoil 2A CTD phosphatase subunit activating transcription factor 7 interacting protein ubiquitin protein ligase E3 component n-recognin 5 arginine-glutamic acid dipeptide repeats arginine-glutamic acid dipeptide repeats pumilio RNA binding family member 2 RUN and cysteine rich domain containing beclin interacting protein RUN and cysteine rich domain containing beclin interacting protein RUN and cysteine rich domain containing beclin interacting protein vestigial like family member 4 ubiquitin-associated protein 2-like Q7TSC upregulation Q7TSC upregulation Q7TSG upregulation Q7TT upregulation Q80TP upregulation Q80TZ upregulation Q80TZ upregulation Q80U upregulation Q80U upregulation Q80U upregulation Q80U upregulation Q80V upregulation Q80X upregulation AHTDEPHS(ph)PQSK elac ribonuclease Z 2 Q80Y upregulation RAHTDEPHS(ph)PQSK elac ribonuclease Z 2 Q80Y upregulation TAS(ph)NAEQYK SMGTGDSAGVEVPSS(p h)plrr LCS(ph)SSSSDTSPR actin filament associated protein zinc finger C3HCtype containing zinc finger C3HCtype containing Q80YS upregulation Q80YV upregulation Q80YV upregulation
20 SQS(ph)LPGADSLLAKPI DK EQMMNSSVSSGSGS(ph) LR transformation/trans cription domain associated protein discs large MAGUK scaffold protein Q80YV upregulation Q8D upregulation SS(ph)PTPHSGPCPSR proline rich 5 Q82A upregulation AEEGGESEGDAS(ph)EK DAK SRTGSES(ph)SQTGASA TSGR TGS(ph)ESSQTGASATS GR TGSESS(ph)QTGASATS GR LIHQFLDEDSDPMLS(ph )PR SRS(ph)QPCVLNDK FGGQPCQGAPGSAPCG QAGDSWS(ph)PDPHPVG GGR SRS(ph)ESETSTMAAK HGAPAAPS(ph)PPPR VVPQQITHTS(ph)PR LS(ph)PEPVAHR SRT(ph)PSASHEEQQE TPS(ph)ASHEEQQE SRTPSAS(ph)HEEQQE VENDENNETLS(ph)EPG ESSKEENCSK SSPGNLRDQS(ph)PK LAGSGDDS(ph)PKRK LKSGMS(ph)PEQSK HSLSGSS(ph)PGMK QSSS(ph)PYEDKDKK REISSS(ph)PTSK HIV- Tat specific factor eukaryotic translation initiation factor 4B eukaryotic translation initiation factor 4B eukaryotic translation initiation factor 4B family with sequence similarity 3, member A family with sequence similarity 53 member B family with sequence similarity 53 member B chromosome 8 open reading frame 25 TBC domain family member 0B Sin3A associated protein 30 Smad nuclear interacting protein small glutamine rich tetratricopeptide repeat containing alpha small glutamine rich tetratricopeptide repeat containing alpha small glutamine rich tetratricopeptide repeat containing alpha SWI5 dependent homologous recombination repair protein protein phosphatase regulatory subunit 8 protein phosphatase regulatory subunit 8 Q8BGC upregulation Q8BGD upregulation Q8BGD upregulation Q8BGD upregulation Q8BGI upregulation Q8BGR upregulation Q8BGR upregulation Q8BH upregulation Q8BHL upregulation Q8BIH upregulation Q8BIZ upregulation Q8BJU upregulation Q8BJU upregulation Q8BJU upregulation Q8BP upregulation Q8BQ upregulation Q8BQ upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation
21 QSHSESS(ph)PDGEVK SRTS(ph)PVTR DGLPRT(ph)PSRR SGS(ph)SQELDGKPSAS PQER SGSS(ph)QELDGKPSAS PQER SRTS(ph)PVSR SVT(ph)PQRER SRS(ph)GSSQELDGKPS ASPQER HSLS(ph)GSSPGMKDTP QTPSR SST(ph)PSSPTGTSSTDS GGQHLGWGEQQGQWL R NGKEQYVPPRS(ph)PK SRS(ph)QPCDLDAR LGS(ph)MDSFER HAS(ph)APSHVQPSDSE K LGS(ph)MDSFER ST(ph)TPVKEKEHSK AADEKGS(ph)PRTEDE GK KAPARPSS(ph)ASATPR FNSDSHS(ph)PK SQCRNSPS(ph)NLSSSSE TGSGGGTYR EYGST(ph)SSIDK LSS(ph)TGGQTPR S(ph)PTKSSLDYR SPLRRS(ph)PPR TNSMSSSGLGS(ph)PNR YTEQERS(ph)PR phosphorylase kinase regulatory subunit alpha 2 La ribonucleoprotein domain family member 4 family with sequence similarity 53 member C TBC domain family member 4 TBC domain family member 4 TBC domain family member 4 splicing regulatory glutamic acid and lysine rich protein splicing regulatory glutamic acid and lysine rich protein microtubule associated protein S Rho GTPase activating protein 2 signal induced proliferation associated like signal induced proliferation associated like IWS, SUPT6H interacting protein RNA binding motif protein 4 RNA binding motif protein 4 CCR4-NOT transcription complex subunit 2 CWC22 spliceosome associated protein homolog Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BTI upregulation Q8BWJ upregulation Q8BWW upregulation Q8BXQ upregulation Q8BYJ upregulation Q8BYJ upregulation Q8BYJ E-08 upregulation Q8BZX upregulation Q8BZX upregulation Q8C upregulation Q8C0D upregulation Q8C0T upregulation Q8C0T upregulation Q8CD upregulation Q8C2Q upregulation Q8C2Q upregulation Q8C5L upregulation Q8C5N upregulation TAS(ph)LNQRPR abl interactor Q8CBW downregulation LGSQHS(ph)PGR abl interactor Q8CBW downregulation
22 HNS(ph)TTSSTSSGGYR R LLLLASS(ph)PTER GRRPS(ph)GPEGAAR ISQRDPS(ph)PESK KGGS(ph)PGLESR SPS(ph)QEPSAPGKAEA VGEQAR KAEGEPQEES(ph)PLKS K LRCDS(ph)ADLR EVQS(ph)PEQVKSEK EVQS(ph)PEQVK QKS(ph)PEIHR S(ph)PAKTITPQNAPR KAEGEPQEES(ph)PLK RYSSSGT(ph)PSSASPAL SR abl interactor Q8CBW upregulation Rho GTPase activating protein 29 polyhomeotic homolog 3 polyhomeotic homolog 3 polyhomeotic homolog 3 eukaryotic translation initiation factor 3 subunit B BCL2 associated transcription factor BCL2 associated transcription factor BCL2 associated transcription factor BCL2 associated transcription factor BCL2 associated transcription factor BCL2 associated transcription factor BCL2 associated transcription factor nuclear factor of activated T cells 4 Q8CGF upregulation Q8CHP upregulation Q8CHP upregulation Q8CHP upregulation Q8JZQ upregulation Q8K upregulation Q8K upregulation Q8K upregulation Q8K upregulation Q8K upregulation Q8K upregulation Q8K upregulation Q8K upregulation ELPSAGS(ph)RDK dynamin like Q8KM upregulation STQSENQHQGAQDTSD LMS(ph)PSKR EKTMSS(ph)DDEECSAK integrator complex subunit 0 integrator complex subunit 0 Q8K2A upregulation Q8K2A upregulation SYS(ph)PDGKESPSDK matrin 3 Q8K upregulation SYS(ph)PDGKESPSDKK matrin 3 Q8K upregulation GT(ph)VTGERQSGDGQ ESTEPVENKVGK GTVTGERQS(ph)GDGQ ESTEPVENK DAPRPDHPPHDGHS(p h)pasr cancer susceptibility 3 cancer susceptibility 3 WD repeat domain 33 Q8K3W upregulation Q8K3W upregulation Q8K4P upregulation HDLDAS(ph)PPR BUD3 homolog Q8R upregulation HDLDAS(ph)PPRK BUD3 homolog Q8R upregulation S(ph)MSIDDTPR KPASVSPTTPTS(ph)PTE GEAS SS(ph)PVHIIATSK S(ph)PGPPALKHPTSK RKAS(ph)PEPEGETAGK VEEEQEADEEDVS(ph)E EEAEDREGASK GTGDCS(ph)DEEVDGK ADGADAK dedicator of cytokinesis 7 Q8RA upregulation dynein cytoplasmic Q8RQ upregulation light intermediate chain HMG-box Q8R upregulation transcription factor interferon regulatory Q8R3Y upregulation factor 2 binding protein interferon regulatory Q8R3Y upregulation factor 2 binding protein thioredoxin related Q8VBT upregulation transmembrane protein myosin heavy chain 9 Q8VDD upregulation
RelEx v. 1.0 (2004) Michael J. MacCoss and John D. Venable, The Scripps Research Institute, La Jolla, CA (2002)
 RelEx v. 1.0 (2004) Michael J. MacCoss and John D. Venable, The Scripps Research Institute, La Jolla, CA (2002) P CE00015 locus:drs-1 Aspartyl-tRNA synthetase status:partially_confirm 0.876 0.074 7 R U
RelEx v. 1.0 (2004) Michael J. MacCoss and John D. Venable, The Scripps Research Institute, La Jolla, CA (2002) P CE00015 locus:drs-1 Aspartyl-tRNA synthetase status:partially_confirm 0.876 0.074 7 R U
Supporting Information for Proteomics DOI /pmic
 Supporting Information for Proteomics DOI 10.1002/pmic.200401063 Linda L. Manza, Sheryl L. Stamer, Amy-Joan L. Ham, Simona G. Codreanu and Daniel C. Liebler Sample preparation and digestion for proteomic
Supporting Information for Proteomics DOI 10.1002/pmic.200401063 Linda L. Manza, Sheryl L. Stamer, Amy-Joan L. Ham, Simona G. Codreanu and Daniel C. Liebler Sample preparation and digestion for proteomic
Table 1s. Identified proteins along with modifications. Δ m/z Meta Range Sequence Modifications Accession Protein
 Table 1s. Identified proteins along with modifications Δ m/z Meta Range Sequence Modifications Accession Protein [ppm] Score -28.15 43.0 1256 - R.TLEDQVNELK.S 1265 6.52 27.5 11-32 R.AIQANINIPMGAFRPGAGQPPR.R
Table 1s. Identified proteins along with modifications Δ m/z Meta Range Sequence Modifications Accession Protein [ppm] Score -28.15 43.0 1256 - R.TLEDQVNELK.S 1265 6.52 27.5 11-32 R.AIQANINIPMGAFRPGAGQPPR.R
Supporting Information
 Supporting Information Expanding the dipeptidyl peptidase 4-regulated peptidome via an optimized peptidomics platform Arthur D. Tinoco, Debarati M. Tagore, and Alan Saghatelian* Department of Chemistry
Supporting Information Expanding the dipeptidyl peptidase 4-regulated peptidome via an optimized peptidomics platform Arthur D. Tinoco, Debarati M. Tagore, and Alan Saghatelian* Department of Chemistry
Regulering av DNA Transkripsjon i Eukaryote Organismer. ID, Kull 99, Vår 2001 Frank Skorpen IKM, DMF
 Regulering av DNA Transkripsjon i Eukaryote Organismer ID, Kull 99, Vår 2001 Frank Skorpen IKM, DMF 1 Regulering av gen-uttrykk på mange nivåer Klargjøring av DNA Transkripsjon Initiering Stopp hnrna prosessering
Regulering av DNA Transkripsjon i Eukaryote Organismer ID, Kull 99, Vår 2001 Frank Skorpen IKM, DMF 1 Regulering av gen-uttrykk på mange nivåer Klargjøring av DNA Transkripsjon Initiering Stopp hnrna prosessering
HPV og molekylærbiologi Molekylære mekanismer bak HPV-indusert kreftutvikling
 HPV og molekylærbiologi Molekylære mekanismer bak HPV-indusert kreftutvikling Irene Kraus Christiansen Nasjonalt referanselaboratorium for HPV Molekylære mekanismer ved kreftutvikling Forståelse av samspillet
HPV og molekylærbiologi Molekylære mekanismer bak HPV-indusert kreftutvikling Irene Kraus Christiansen Nasjonalt referanselaboratorium for HPV Molekylære mekanismer ved kreftutvikling Forståelse av samspillet
SERK1/2 Acts as a Partner of EMS1 to Control Anther Cell Fate Determination in Arabidopsis
 SERK1/2 Acts as a Partner of EMS1 to Control Anther Cell Fate Determination in Arabidopsis Supplemental Data Supplemental Figure S1 Supplemental Figure S1. Identification of the ems1-2 weak allele. A,
SERK1/2 Acts as a Partner of EMS1 to Control Anther Cell Fate Determination in Arabidopsis Supplemental Data Supplemental Figure S1 Supplemental Figure S1. Identification of the ems1-2 weak allele. A,
EKSAMEN I EMNE TBT4102 BIOKJEMI I. 3. desember 2012 kl
 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR BIOTEKNOLOGI Faglig kontakt under eksamen: Institutt for bioteknologi, Gløshaugen Kjell M. Vårum, tlf. 93022165 EKSAMEN I EMNE TBT4102 BIOKJEMI
NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR BIOTEKNOLOGI Faglig kontakt under eksamen: Institutt for bioteknologi, Gløshaugen Kjell M. Vårum, tlf. 93022165 EKSAMEN I EMNE TBT4102 BIOKJEMI
REGULERING AV TRANSKRIPSJON I EUKARYOTE ORGANISMER
 REGULERING AV TRANSKRIPSJON I EUKARYOTE ORGANISMER av Frank Skorpen, IKM, DMF, NTNU FORELESNING April 2001 Cellebiologi - Hovedfag 1 REGULERING AV TRANSKRIPSJON I EUKARYOTE ORGANISMER Det er flere nøkkel-mekanismer
REGULERING AV TRANSKRIPSJON I EUKARYOTE ORGANISMER av Frank Skorpen, IKM, DMF, NTNU FORELESNING April 2001 Cellebiologi - Hovedfag 1 REGULERING AV TRANSKRIPSJON I EUKARYOTE ORGANISMER Det er flere nøkkel-mekanismer
EKSAMEN I EMNE TBT4100 BIOKJEMI GRUNNKURS Tirsdag 14. desember 2006 Tid: kl. 09:00 14:00
 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR BIOTEKNOLOGI 1 Faglig kontakt under ekasmen: Institutt for bioteknologi, Gløshaugen Førsteamanuensis Sergey B. Zotchev, tlf. 98679. EKSAMEN
NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR BIOTEKNOLOGI 1 Faglig kontakt under ekasmen: Institutt for bioteknologi, Gløshaugen Førsteamanuensis Sergey B. Zotchev, tlf. 98679. EKSAMEN
Eksamensoppgave i Bi3016 Celle og molekylærbiologi Examination in Bi3016 Cell and molecular biology
 Institutt for biologi / Department of Biology Eksamensoppgave i Bi3016 Celle og molekylærbiologi Examination in Bi3016 Cell and molecular biology Faglig kontakt under eksamen / Contact person during exam:
Institutt for biologi / Department of Biology Eksamensoppgave i Bi3016 Celle og molekylærbiologi Examination in Bi3016 Cell and molecular biology Faglig kontakt under eksamen / Contact person during exam:
Oncogenic Mutations Affecting Cell Proliferation
 Oncogenic Mutations Affecting Cell Proliferation Fra RTK til Nucleus (Boka s.1070-74) Normalt kreves et vekst stimulerende signal ( growth factor eks. PDGF, EGF, NGF) for at celler skal gå inn i celledeling,
Oncogenic Mutations Affecting Cell Proliferation Fra RTK til Nucleus (Boka s.1070-74) Normalt kreves et vekst stimulerende signal ( growth factor eks. PDGF, EGF, NGF) for at celler skal gå inn i celledeling,
Hvordan holde styr på proteasene?? Harald; 31.01.08
 Hvordan holde styr på proteasene?? Harald; 31.01.08 Protease? Proteinase? Spalter proteiner Peptidase? Spalter peptider? Spalter peptidbindinger Exo-peptidaser Endo-peptidaser Endo Ekso- Proteaser er enzymer!
Hvordan holde styr på proteasene?? Harald; 31.01.08 Protease? Proteinase? Spalter proteiner Peptidase? Spalter peptider? Spalter peptidbindinger Exo-peptidaser Endo-peptidaser Endo Ekso- Proteaser er enzymer!
FYS 3710 Biofysikk og Medisinsk Fysikk, DNA, RNA, Translasjon, Transkripsjon Proteinsyntese, Cellesyklus
 FYS 3710 Biofysikk og Medisinsk Fysikk, 2017 7 DNA, RNA, Translasjon, Transkripsjon Proteinsyntese, Cellesyklus Einar Sagstuen, Fysisk institutt, UiO 18.09.2017 1 DNA A / C / G / T 2 -deoxyribose monofosfate
FYS 3710 Biofysikk og Medisinsk Fysikk, 2017 7 DNA, RNA, Translasjon, Transkripsjon Proteinsyntese, Cellesyklus Einar Sagstuen, Fysisk institutt, UiO 18.09.2017 1 DNA A / C / G / T 2 -deoxyribose monofosfate
Dynamic Programming Longest Common Subsequence. Class 27
 Dynamic Programming Longest Common Subsequence Class 27 Protein a protein is a complex molecule composed of long single-strand chains of amino acid molecules there are 20 amino acids that make up proteins
Dynamic Programming Longest Common Subsequence Class 27 Protein a protein is a complex molecule composed of long single-strand chains of amino acid molecules there are 20 amino acids that make up proteins
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION IKK phosphorylation regulates RPS3 nuclear translocation and NF- B function during Escherichia coli O157:H7 infection Fengyi Wan 1,2, Amanda Weaver 1, Xiaofei Gao 3, Michael Bern
SUPPLEMENTARY INFORMATION IKK phosphorylation regulates RPS3 nuclear translocation and NF- B function during Escherichia coli O157:H7 infection Fengyi Wan 1,2, Amanda Weaver 1, Xiaofei Gao 3, Michael Bern
Cellesyklus. Medisin stadium IA, 17. september 2012
 Cellesyklus Medisin stadium IA, 17. september 2012 Trude Helen Flo Cellesyklus: En oversikt Definisjoner De ulike fasene av cellesyklus Regulering av cellesyklus Kort om apoptose Kort om stamceller Cellesyklus:
Cellesyklus Medisin stadium IA, 17. september 2012 Trude Helen Flo Cellesyklus: En oversikt Definisjoner De ulike fasene av cellesyklus Regulering av cellesyklus Kort om apoptose Kort om stamceller Cellesyklus:
GENER, genregulering, og genfamilier
 GENER, genregulering, og genfamilier 1-A, H-11 Forelesning 21.11.11 Frank Skorpen, Institutt for Laboratoriemedisin, Barne- og Kvinnesykdommer, DMF, NTNU Gener Kromosom, kromatin og DNA Hva er et gen?
GENER, genregulering, og genfamilier 1-A, H-11 Forelesning 21.11.11 Frank Skorpen, Institutt for Laboratoriemedisin, Barne- og Kvinnesykdommer, DMF, NTNU Gener Kromosom, kromatin og DNA Hva er et gen?
Editor-in-Chief. Associate Editor. Editorial Board. Michael J. Williams University of Aberdeen. Klemens Rottner University of Bonn
 Editor-in-Chief Michael J. Williams University of Aberdeen Associate Editor Klemens Rottner University of Bonn Udo Häcker Lund University Johanna Ivaska VTT Medical Biotechnology University of Turku Ingo
Editor-in-Chief Michael J. Williams University of Aberdeen Associate Editor Klemens Rottner University of Bonn Udo Häcker Lund University Johanna Ivaska VTT Medical Biotechnology University of Turku Ingo
Supplementary Materials for
 stm.sciencemag.org/cgi/content/full/11/56/eaau8217/dc1 Supplementary Materials for Treating murine inflammatory diseases with an anti-erythrocyte antibody Andrew R. Crow, Rick Kapur, Sandra Koernig, Ian
stm.sciencemag.org/cgi/content/full/11/56/eaau8217/dc1 Supplementary Materials for Treating murine inflammatory diseases with an anti-erythrocyte antibody Andrew R. Crow, Rick Kapur, Sandra Koernig, Ian
UNIVERSITETET I OSLO
 UNIVERSITETET I OSLO Side 1 Det matematisk-naturvitenskapelige fakultet Bokmål Eksamen i MBV 3010, Videregående cellebiologi. Eksamensdag: august 2006. Tid for eksamen: kl (3 timer). Oppgavesettet er på
UNIVERSITETET I OSLO Side 1 Det matematisk-naturvitenskapelige fakultet Bokmål Eksamen i MBV 3010, Videregående cellebiologi. Eksamensdag: august 2006. Tid for eksamen: kl (3 timer). Oppgavesettet er på
Hva viser genanalyser av muskulatur hos laks med mørke flekker. Aleksei Krasnov, Hooman Moghadam Nofima, Ås
 Hva viser genanalyser av muskulatur hos laks med mørke flekker Aleksei Krasnov, Hooman Moghadam Nofima, Ås Spørsmål Har mørke flekker lik eller ulik profil? Vevsskade og nydannelse av vev? Type betennelse?
Hva viser genanalyser av muskulatur hos laks med mørke flekker Aleksei Krasnov, Hooman Moghadam Nofima, Ås Spørsmål Har mørke flekker lik eller ulik profil? Vevsskade og nydannelse av vev? Type betennelse?
NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE TFY4260 CELLEBIOLOGI OG CELLULÆR BIOFYSIKK
 Side av 1 av 4 Studentnr... Studieprogram... Sidenr. NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE TFY4260 CELLEBIOLOGI OG CELLULÆR BIOFYSIKK Faglig kontakt under eksamen:
Side av 1 av 4 Studentnr... Studieprogram... Sidenr. NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE TFY4260 CELLEBIOLOGI OG CELLULÆR BIOFYSIKK Faglig kontakt under eksamen:
Uønsket opptak av store molekyler i lever et problem som kan overkommes?
 Uønsket opptak av store molekyler i lever et problem som kan overkommes? Bård Smedsrød Founder and CSO Blood clearance function of the liver: Fate of i.v. administered macromolecule ( 125 I-FSA) Blood
Uønsket opptak av store molekyler i lever et problem som kan overkommes? Bård Smedsrød Founder and CSO Blood clearance function of the liver: Fate of i.v. administered macromolecule ( 125 I-FSA) Blood
UNIVERSITETET I OSLO
 UNIVERSITETET I OSLO Side 1 Det matematisk-naturvitenskapelige fakultet Eksamen i MBV 3010, Videregående cellebiologi. Eksamensdag: 14. juni 2006. Tid for eksamen: kl 14.30 (3 timer). Oppgavesettet er
UNIVERSITETET I OSLO Side 1 Det matematisk-naturvitenskapelige fakultet Eksamen i MBV 3010, Videregående cellebiologi. Eksamensdag: 14. juni 2006. Tid for eksamen: kl 14.30 (3 timer). Oppgavesettet er
Exercise 1: Phase Splitter DC Operation
 Exercise 1: DC Operation When you have completed this exercise, you will be able to measure dc operating voltages and currents by using a typical transistor phase splitter circuit. You will verify your
Exercise 1: DC Operation When you have completed this exercise, you will be able to measure dc operating voltages and currents by using a typical transistor phase splitter circuit. You will verify your
!"" #$ % <'/ & ' & & " E*.E *N 9 " 9 ) $ 9 ' &" )*./W BN 9 '" 9E * )* * 9 '" \./W 45 J = [\ T [\ > NO 1Z % H & 9: TG 23 Y*[\ $ * '
 !"" #$ %1 21+ 3 1 NO 1Z % H & 9: TG 23 Y*[\ $ * ' =N> Y* TG *! > " 9: 23J #$%&' F '3 * (23 )* +0,-G.0XO/0
!"" #$ %1 21+ 3 1 NO 1Z % H & 9: TG 23 Y*[\ $ * ' =N> Y* TG *! > " 9: 23J #$%&' F '3 * (23 )* +0,-G.0XO/0
FYS 3710 Biofysikk og Medisinsk Fysikk, Aminosyrer, Polypeptider, Proteiner
 FYS 3710 Biofysikk og Medisinsk Fysikk, 2016 5 Aminosyrer, Polypeptider, Proteiner Einar Sagstuen, Fysisk institutt, UiO 06.09.2016 1 sp n -hybridisering: for hovedkvantetall N=2 er de fire valensorbitalene
FYS 3710 Biofysikk og Medisinsk Fysikk, 2016 5 Aminosyrer, Polypeptider, Proteiner Einar Sagstuen, Fysisk institutt, UiO 06.09.2016 1 sp n -hybridisering: for hovedkvantetall N=2 er de fire valensorbitalene
Skeleton Development. Patricia Ducy HHSC1616 x
 Skeleton Development Patricia Ducy HHSC1616 x5-9299 pd2193@columbia.edu 1 Skeleton 200 elements Two tissues: cartilage, bone Three cell types: chondrocytes, osteoblasts, osteoclasts Growth Formation Resorption
Skeleton Development Patricia Ducy HHSC1616 x5-9299 pd2193@columbia.edu 1 Skeleton 200 elements Two tissues: cartilage, bone Three cell types: chondrocytes, osteoblasts, osteoclasts Growth Formation Resorption
Semmelweis University. Genetic and epigenetic regulation of. Dept. GCI. the immune response. András Falus. November
 Genetic and epigenetic regulation of the immune response András Falus Do-Re-Mi November 5. 2013. Semmelweis University Dept. GCI Encyclopedy of DNA elements of human genome released: April, 2012 The human
Genetic and epigenetic regulation of the immune response András Falus Do-Re-Mi November 5. 2013. Semmelweis University Dept. GCI Encyclopedy of DNA elements of human genome released: April, 2012 The human
A study of the contribution of the PHD finger
 Cooperation between the bromodomain and PHD finger of p300 in nucleosome interaction A study of the contribution of the PHD finger Kamilla Breen Thesis submitted in partial fulfilment of the requirements
Cooperation between the bromodomain and PHD finger of p300 in nucleosome interaction A study of the contribution of the PHD finger Kamilla Breen Thesis submitted in partial fulfilment of the requirements
Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
 Side 1 av 12 Norges teknisknaturvitenskapelige universitet Fakultet for naturvitenskap og teknologi Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
Side 1 av 12 Norges teknisknaturvitenskapelige universitet Fakultet for naturvitenskap og teknologi Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
Nucleic Acid Research Group Study:
 Nucleic Acid Research Group 2007-2008 Study: A Comparison of Different Priming Strategies for cdna Synthesis by Reverse Transcriptase as Measured by Real-Time RT-qPCR NARG Survey Results for Types of Primers
Nucleic Acid Research Group 2007-2008 Study: A Comparison of Different Priming Strategies for cdna Synthesis by Reverse Transcriptase as Measured by Real-Time RT-qPCR NARG Survey Results for Types of Primers
Generation of 32 P-labeled MEK, ERK and. Kendrick Labs Inc, Madison, WI
 Generation of 32 P-labeled MEK, ERK and VEGF-R protein standards for 2D gels Jon Johansen Matt Hoelter & Nancy Kendrick* Jon Johansen, Matt Hoelter & Nancy Kendrick Kendrick Labs Inc, Madison, WI www.kendricklabs.com
Generation of 32 P-labeled MEK, ERK and VEGF-R protein standards for 2D gels Jon Johansen Matt Hoelter & Nancy Kendrick* Jon Johansen, Matt Hoelter & Nancy Kendrick Kendrick Labs Inc, Madison, WI www.kendricklabs.com
Besvarelse eksamen i emnet TFY4260 Cellebiologi og cellulær biofysikk 1 juni 2010
 1 Besvarelse eksamen i emnet TFY4260 Cellebiologi og cellulær biofysikk 1 juni 2010 Oppgave 1: Transport over membraner. Na + /K + -ATPase pumpe Na+/K+ pumpen er et transmembranprotein som har 3 bindingssteder
1 Besvarelse eksamen i emnet TFY4260 Cellebiologi og cellulær biofysikk 1 juni 2010 Oppgave 1: Transport over membraner. Na + /K + -ATPase pumpe Na+/K+ pumpen er et transmembranprotein som har 3 bindingssteder
RF Power Capacitors Class1. 5kV Discs
 RF Power Capacitors Class 5kV Discs T H E C E R A M C E X P E R T S RF Power Capacitors Class 5kV Discs The CeramTec Group is a world leader in the design and manufacture of complex electronic ceramic
RF Power Capacitors Class 5kV Discs T H E C E R A M C E X P E R T S RF Power Capacitors Class 5kV Discs The CeramTec Group is a world leader in the design and manufacture of complex electronic ceramic
Det humane genomet. (genom = arvemasse) 3 x 10 9 basepar. ca. 30.000 gen. ferdigsekvensert ca. år 2001
 Det humane genomet (genom = arvemasse) 3 x 10 9 basepar ca. 30.000 gen ferdigsekvensert ca. år 2001 kromosom 21 127 kjente gen 98 ukjente gen Biologi cellekultur organ organismar kva gen er involvert i
Det humane genomet (genom = arvemasse) 3 x 10 9 basepar ca. 30.000 gen ferdigsekvensert ca. år 2001 kromosom 21 127 kjente gen 98 ukjente gen Biologi cellekultur organ organismar kva gen er involvert i
Cytoskjelettet. plasmamembran. Terje Espevik IKM. plasmamembran. Oversikt over Aktinfilamenter Mikrotubuli Intermediærfilamenter
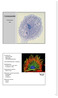 plasmamembran Cytoskjelettet Terje Espevik IKM plasmamembran Oversikt over Aktinfilamenter Mikrotubuli Intermediærfilamenter Intermediærfilamenter; inndeling Aktinfilamenter Struktur av G- og F-aktin Funksjoner
plasmamembran Cytoskjelettet Terje Espevik IKM plasmamembran Oversikt over Aktinfilamenter Mikrotubuli Intermediærfilamenter Intermediærfilamenter; inndeling Aktinfilamenter Struktur av G- og F-aktin Funksjoner
Exosomes Mediate Epithelium-Mesenchyme Crosstalk in Organ Development
 1 Exosomes Mediate Epithelium-Mesenchyme Crosstalk in Organ Development Nan Jiang 1,2, Lusai Xiang 2,5, Ling He 2,5, Guodong Yang 2, Jinxuan Zheng 2,5, Chenglin Wang 2,6, Yimei Zhang 3, Sainan Wang 2,
1 Exosomes Mediate Epithelium-Mesenchyme Crosstalk in Organ Development Nan Jiang 1,2, Lusai Xiang 2,5, Ling He 2,5, Guodong Yang 2, Jinxuan Zheng 2,5, Chenglin Wang 2,6, Yimei Zhang 3, Sainan Wang 2,
Amplifikasjonsteknikker - andre metoder
 Amplifikasjonsteknikker - andre metoder Svein Arne Nordbø TH-28973 17.03.15 Alternative amplifikasjonsmetoder Templat-amplifikasjons metoder Signal-amplifikasjonsmetoder Templat-amplifikasjons metoder
Amplifikasjonsteknikker - andre metoder Svein Arne Nordbø TH-28973 17.03.15 Alternative amplifikasjonsmetoder Templat-amplifikasjons metoder Signal-amplifikasjonsmetoder Templat-amplifikasjons metoder
RF Power Capacitors Class kV Discs with Moisture Protection
 RF Power Capacitors Class 0-20kV Discs with Moisture Protection T H E C E R A M I C E X P E R T S RF Power Capacitors Class 0-20kV Discs with Moisture Protection The CeramTec Group is a world leader in
RF Power Capacitors Class 0-20kV Discs with Moisture Protection T H E C E R A M I C E X P E R T S RF Power Capacitors Class 0-20kV Discs with Moisture Protection The CeramTec Group is a world leader in
Examination paper for Bi2014 Molecular Biology
 Department of Biology Examination paper for Bi2014 Molecular Biology Academic contact during examination: Professor Atle M. Bones Phone: 91897237 (73598692) Examination date/eksamensdag: 7. December 2016
Department of Biology Examination paper for Bi2014 Molecular Biology Academic contact during examination: Professor Atle M. Bones Phone: 91897237 (73598692) Examination date/eksamensdag: 7. December 2016
Myelomatose -sett fra et cellebiologisk ståsted
 Myelomatose -sett fra et cellebiologisk ståsted KG Jebsen Center for Myeloma Research Department of Cancer Research and Molecular Medicine - NTNU anders.sundan@ntnu.no 1 Multiple Myeloma ( Myelomatose
Myelomatose -sett fra et cellebiologisk ståsted KG Jebsen Center for Myeloma Research Department of Cancer Research and Molecular Medicine - NTNU anders.sundan@ntnu.no 1 Multiple Myeloma ( Myelomatose
NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE TFY4260 CELLEBIOLOGI OG CELLULÆR BIOFYSIKK
 Side av 1 av5 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE TFY4260 CELLEBIOLOGI OG CELLULÆR BIOFYSIKK Faglig kontakt under eksamen: Catharina Davies Tel 73593688 eller
Side av 1 av5 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE TFY4260 CELLEBIOLOGI OG CELLULÆR BIOFYSIKK Faglig kontakt under eksamen: Catharina Davies Tel 73593688 eller
Cellebiologiske prosesser som respons på virusinfeksjon
 Cellebiologiske prosesser som respons på virusinfeksjon PBM 336 2005 Siri Mjaaland Infeksjoner - immunresponser 1 Figure 2-49 Interferoner Uspesifikk immunitet viral infeksjon stimulerer direkte produksjon
Cellebiologiske prosesser som respons på virusinfeksjon PBM 336 2005 Siri Mjaaland Infeksjoner - immunresponser 1 Figure 2-49 Interferoner Uspesifikk immunitet viral infeksjon stimulerer direkte produksjon
Forelesninger i BI Cellebiologi. Denaturering og renaturering. Figure 3-13
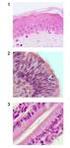 Figure 3.9 Denaturering og renaturering Figure 3-13 Denaturering og renaturering Figure 3-14 Viser tre trinn i refolding av et protein som har vært denaturert. Molten globule -formen er en intermediær
Figure 3.9 Denaturering og renaturering Figure 3-13 Denaturering og renaturering Figure 3-14 Viser tre trinn i refolding av et protein som har vært denaturert. Molten globule -formen er en intermediær
Cellesignalisering II: Reseptor tyrosin kinaser, cytosoliske kinaser
 Cellesignalisering II: Reseptor tyrosin kinaser, cytosoliske kinaser! Introduksjon! Definisjon og klassifisering! Kinasefamilier: Receptor/cytosol! Receptor Tyrosin kinase-mediert signalisering! MAP kinase
Cellesignalisering II: Reseptor tyrosin kinaser, cytosoliske kinaser! Introduksjon! Definisjon og klassifisering! Kinasefamilier: Receptor/cytosol! Receptor Tyrosin kinase-mediert signalisering! MAP kinase
Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
 Side 1 av 13 Norges teknisknaturvitenskapelige universitet Fakultet for naturvitenskap og teknologi Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
Side 1 av 13 Norges teknisknaturvitenskapelige universitet Fakultet for naturvitenskap og teknologi Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
Anne Helene Barsnes 03.06.2011
 TBT4170 Terms Begreper og biologiske prosesser i Bioteknologi Anne Helene Barsnes 03.06.2011 Innholdsfortegnelse 1. Tree of life/cell biology/basic chemistry of life... 4 Livets tre... 4 2. Enzymes...
TBT4170 Terms Begreper og biologiske prosesser i Bioteknologi Anne Helene Barsnes 03.06.2011 Innholdsfortegnelse 1. Tree of life/cell biology/basic chemistry of life... 4 Livets tre... 4 2. Enzymes...
Fastere filet - Aktivitet 2
 Fastere filet - Aktivitet 2 MRI, protease og protein analyser Iciar Martinez 1,2, Rasa Slizyté 1, Pål Anders Wang 2,Stine W. Dahle 1, Michiaki Yamashita 3, Ragnar Olsen 2, Emil Veliuylin 1 og Ulf Erikson
Fastere filet - Aktivitet 2 MRI, protease og protein analyser Iciar Martinez 1,2, Rasa Slizyté 1, Pål Anders Wang 2,Stine W. Dahle 1, Michiaki Yamashita 3, Ragnar Olsen 2, Emil Veliuylin 1 og Ulf Erikson
Foreleser: Eivind Coward, kontor 5. etg. Datablokken. coward@ii.uib.no Gruppeleder: Harald Barsnes
 Foreleser: Eivind Coward, kontor 5. etg. Datablokken. coward@ii.uib.no Gruppeleder: Harald Barsnes Forelesninger: tirsdag og fredag 12 14 rom 2104 Øvinger: fredag 10 12 rom 2143 Gi en innføring i noen
Foreleser: Eivind Coward, kontor 5. etg. Datablokken. coward@ii.uib.no Gruppeleder: Harald Barsnes Forelesninger: tirsdag og fredag 12 14 rom 2104 Øvinger: fredag 10 12 rom 2143 Gi en innføring i noen
Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
 Side 1 av 13 Norges teknisknaturvitenskapelige universitet Fakultet for naturvitenskap og teknologi Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
Side 1 av 13 Norges teknisknaturvitenskapelige universitet Fakultet for naturvitenskap og teknologi Institutt for biologi Faglig kontaktperson under eksamen: Berit Johansen (98691), Janne Østvang (51278)
Modeling the immune system
 Master Modeling the immune system nnate mmunity Innate immunity or natural immunity refers to body's defense mechanisms that: are available instantaneously and for the first few day after microbial infection.
Master Modeling the immune system nnate mmunity Innate immunity or natural immunity refers to body's defense mechanisms that: are available instantaneously and for the first few day after microbial infection.
Ti grunner for å leve i håpet Dr Ed Wild
 Ti grunner for å leve i håpet Dr Ed Wild Clinical Lecturer, UCL Institute of Neurology Specialist Registrar, National Hospital for Neurology & Neurosurgery Co-founder & Editor-in-Chief, HDBuzz Huntington
Ti grunner for å leve i håpet Dr Ed Wild Clinical Lecturer, UCL Institute of Neurology Specialist Registrar, National Hospital for Neurology & Neurosurgery Co-founder & Editor-in-Chief, HDBuzz Huntington
Supplemental Information
 Supplemental Information Figure S1. 2D gel electrophoresis of ipt-transgenic broccoli at harvesting and after cooking. Protein composition of ipt-transgenic broccoli lines 102 and 103 and non-transgenic
Supplemental Information Figure S1. 2D gel electrophoresis of ipt-transgenic broccoli at harvesting and after cooking. Protein composition of ipt-transgenic broccoli lines 102 and 103 and non-transgenic
Figure S1. Figure S2.
 Figure S1. MALS-SEC trace of Tm14S receptor fragment (residues 107-192). Rayleigh ratio (LS) and differential refractive index (RI) shown as a function of retention time. Due to the lack of Trp and Tyr
Figure S1. MALS-SEC trace of Tm14S receptor fragment (residues 107-192). Rayleigh ratio (LS) and differential refractive index (RI) shown as a function of retention time. Due to the lack of Trp and Tyr
Forelesninger i BI Cellebiologi. Enzymer : senker aktiveringsenergien. Figure 6.13
 Enzymer : senker aktiveringsenergien Figure 6.13 Aktive seter : camp-avhengig protein kinase *For å illustrere hvordan det aktive setet binder et spesifikt substrat er valgt som eksempel camp-avhengig
Enzymer : senker aktiveringsenergien Figure 6.13 Aktive seter : camp-avhengig protein kinase *For å illustrere hvordan det aktive setet binder et spesifikt substrat er valgt som eksempel camp-avhengig
RF Power Capacitors Class , 20 & 30 mm Barrel Transmitting Types
 RF Power Capacitors Class 2.7, 20 & 30 mm Barrel Transmitting Types T H E C E R A M I C E X P E R T S RF Power Capacitors Class 2.7, 20 & 30 mm Barrel Transmitting Types The CeramTec Group is a world leader
RF Power Capacitors Class 2.7, 20 & 30 mm Barrel Transmitting Types T H E C E R A M I C E X P E R T S RF Power Capacitors Class 2.7, 20 & 30 mm Barrel Transmitting Types The CeramTec Group is a world leader
Kapittel 14: Det eukaryote genom og dets uttrykksregulering
 Kapittel 14: Det eukaryote genom og dets uttrykksregulering Innhold: 1. Det humane genom 2. Struktur av protein-kodende gener 3. RNA processering 4. Transkripsjonell kontroll 5. Posttranskripsjonell kontroll
Kapittel 14: Det eukaryote genom og dets uttrykksregulering Innhold: 1. Det humane genom 2. Struktur av protein-kodende gener 3. RNA processering 4. Transkripsjonell kontroll 5. Posttranskripsjonell kontroll
UNIVERSITETET I OSLO
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Eksamen i: KJB 492 Bioinformatikk Eksamensdag: Fredag 14. desember 2001 Tid for eksamen: Kl.: 9.00 13.00 Oppgavesettet er på 7 sider. Vedlegg:
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Eksamen i: KJB 492 Bioinformatikk Eksamensdag: Fredag 14. desember 2001 Tid for eksamen: Kl.: 9.00 13.00 Oppgavesettet er på 7 sider. Vedlegg:
BI 212- Protein Sorting - Kap. 17 Syntese og mål for mitokondrie- og kloroplast-proteiner (forts.)
 Syntese og mål for mitokondrie- og kloroplast-proteiner (forts.) Veiene for opptak fra cytosol av kloroplast-proteiner Opptak av proteiner fra cytosol til kloroplaster ligner mye på mitokondrie-importen
Syntese og mål for mitokondrie- og kloroplast-proteiner (forts.) Veiene for opptak fra cytosol av kloroplast-proteiner Opptak av proteiner fra cytosol til kloroplaster ligner mye på mitokondrie-importen
Supplemental Data. Yamamoto et al. (2011). Plant Cell /tpc NADH. e - NAD + Nqo5. Nqo4 Nqo6. Nqo11. Nqo10 Nqo7.
 A NADH Cytoplasm e - Nqo FMN Nqo2 Nqo3 Nqo5 Nqo9 Nqo5 Nqo4 Nqo6 NAD + Membrane Nqo8 Nqo Nqo Nqo7 Nqo4 Nqo3 Nqo2 Periplasm B e - Stroma 3-kD complex? Thylakoid Lumen Fd CRR3 CRRJ CRRL M,N,O H,I,J,K A,B,C,D
A NADH Cytoplasm e - Nqo FMN Nqo2 Nqo3 Nqo5 Nqo9 Nqo5 Nqo4 Nqo6 NAD + Membrane Nqo8 Nqo Nqo Nqo7 Nqo4 Nqo3 Nqo2 Periplasm B e - Stroma 3-kD complex? Thylakoid Lumen Fd CRR3 CRRJ CRRL M,N,O H,I,J,K A,B,C,D
Case 1:11-cr RNS Document 781 Entered on FLSD Docket 03/27/2013 Page 1 of M a u u - g u 'a M M M u..a u i < < < < < < < < <.Q? <.t!
 Cas :2033RNS Dun 78 End n FLSD Dk 03/27/203 Pag f 6 i I jj @ :j j j C I i!, I I! l I : I l!! I ;, ;!, ; 4 k! @ j j ; ;, I I, jji l i I! I j I; l i! l ; : i I I! v z l! l g U U J B g g 6 q; J Y I : 0 ;
Cas :2033RNS Dun 78 End n FLSD Dk 03/27/203 Pag f 6 i I jj @ :j j j C I i!, I I! l I : I l!! I ;, ;!, ; 4 k! @ j j ; ;, I I, jji l i I! I j I; l i! l ; : i I I! v z l! l g U U J B g g 6 q; J Y I : 0 ;
Rusmiddelbruk. Rusmiddelbruk. Hvorfor bruke et rusmiddel? Etablering og utvikling av avhengighet: Hva skjer i hjernen?
 Rusmiddelbruk Etablering og utvikling av avhengighet: Hva skjer i hjernen? JØRG MØRLAND NASJONALT FOLKEHELSEINSTITUTT 1. En enkelt gang 2. Sjelden, intet fast mønster 3. På fester, i helger etc. 4. Mer
Rusmiddelbruk Etablering og utvikling av avhengighet: Hva skjer i hjernen? JØRG MØRLAND NASJONALT FOLKEHELSEINSTITUTT 1. En enkelt gang 2. Sjelden, intet fast mønster 3. På fester, i helger etc. 4. Mer
Bioteknologi/digital teknikk
 Teknologisprang for utvikling av havrommet Bioteknologi/digital teknikk Stig W. Omholt NTNU Biotechnology: the Confluence of Life Sciences, Mathematical Sciences and Engineering Bioteknologi - definisjon
Teknologisprang for utvikling av havrommet Bioteknologi/digital teknikk Stig W. Omholt NTNU Biotechnology: the Confluence of Life Sciences, Mathematical Sciences and Engineering Bioteknologi - definisjon
Kapittel 20, introduksjon
 Kapittel 20, introduksjon Ekstracellulær signalisering Syntese Frigjøring Transport Forandring av cellulær metabolisme, funksjon, utvikling (trigga av reseptor-signal komplekset) Fjerning av signalet Signalisering
Kapittel 20, introduksjon Ekstracellulær signalisering Syntese Frigjøring Transport Forandring av cellulær metabolisme, funksjon, utvikling (trigga av reseptor-signal komplekset) Fjerning av signalet Signalisering
Europa-Universität Viadrina
 !"#!$% & #' #! ( ))% * +%, -.!!! / 0 1!/ %0 2!!/ 0.!!!/ /! 0 / '3 %0 #$ '! 0 4!""2 " '5 + -#! & %%! ( 6+ * $ '. % & 7 7 8 (8 *& *& *( ** *8, 8 87 - - -! )- % 4!!# &! -! ( - / 9:0 ; ; & * 7 4! + /! ) %
!"#!$% & #' #! ( ))% * +%, -.!!! / 0 1!/ %0 2!!/ 0.!!!/ /! 0 / '3 %0 #$ '! 0 4!""2 " '5 + -#! & %%! ( 6+ * $ '. % & 7 7 8 (8 *& *& *( ** *8, 8 87 - - -! )- % 4!!# &! -! ( - / 9:0 ; ; & * 7 4! + /! ) %
Supplementary Figure 2 A
 S U P P L E M E N TA R Y I N F O R M AT I O N DOI:.38/ncb363 Supplementary Figure Mean Residue Elipticity (deg cm / dmol) *^-3-5 7HPSHUDWXUH Û& 4 6 8 - -5 - CC melting -5 CC cooling -3 CCB cooling CCB
S U P P L E M E N TA R Y I N F O R M AT I O N DOI:.38/ncb363 Supplementary Figure Mean Residue Elipticity (deg cm / dmol) *^-3-5 7HPSHUDWXUH Û& 4 6 8 - -5 - CC melting -5 CC cooling -3 CCB cooling CCB
HIV / AIDS -infeksjon - behandling
 HIV / AIDS -infeksjon - behandling HIV / AIDS 1981 første gang anerkjent som distinkt sykdom Opprinnelig overført fra sjimpanse Viruset kan ha sirkulert fra 1915-1941 Trolig sirkulert blant populasjoner
HIV / AIDS -infeksjon - behandling HIV / AIDS 1981 første gang anerkjent som distinkt sykdom Opprinnelig overført fra sjimpanse Viruset kan ha sirkulert fra 1915-1941 Trolig sirkulert blant populasjoner
Proteiner og aminosyrer
 Proteiner og aminosyrer Presentasjonsplan 1/2 Cellen Grunnleggende komponenter DNA til mrna til proteiner Den genetiske koden: Hva er et codon? Presentasjonsplan 2/2 Aminosyrer del 1 Hvilke molekyler er
Proteiner og aminosyrer Presentasjonsplan 1/2 Cellen Grunnleggende komponenter DNA til mrna til proteiner Den genetiske koden: Hva er et codon? Presentasjonsplan 2/2 Aminosyrer del 1 Hvilke molekyler er
BESVARELSE EKSAMEN SIF4070 CELLEBIOLOGI 9. MAI 2003
 1 BESVARELSE EKSAMEN SIF4070 CELLEBIOLOGI 9. MAI 2003 Oppgave 1 a) Transport over membraner Glukose transporteres inn i leverceller med sin konsentrasjonsgradient og ved hjelp av et bæreprotein, dvs at
1 BESVARELSE EKSAMEN SIF4070 CELLEBIOLOGI 9. MAI 2003 Oppgave 1 a) Transport over membraner Glukose transporteres inn i leverceller med sin konsentrasjonsgradient og ved hjelp av et bæreprotein, dvs at
Fosfolipaser, identifikasjon av intracellulære signaliseringsdomener og integrering av multiple signal
 Fosfolipaser, identifikasjon av intracellulære signaliseringsdomener og integrering av multiple signal! Introduksjon! Intra/inter-cellulære lipid sek. budbringere! Fosfolipase A2! Fosfolipase C og D! Sfingomyelinase!
Fosfolipaser, identifikasjon av intracellulære signaliseringsdomener og integrering av multiple signal! Introduksjon! Intra/inter-cellulære lipid sek. budbringere! Fosfolipase A2! Fosfolipase C og D! Sfingomyelinase!
NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE SIF4070 CELLEBIOLOGI
 Side 1 av 6 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Faglig kontakt under eksamen: professor Catharina Davies Tel.73593688 Eksamensdato: 9.
Side 1 av 6 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Faglig kontakt under eksamen: professor Catharina Davies Tel.73593688 Eksamensdato: 9.
Introduksjon til Biokjemi. Ingar Leiros, Institutt for Kjemi, UiT
 Introduksjon til Biokjemi Ingar Leiros, Institutt for Kjemi, UiT Biokjemi Biokjemi (Wikipedia): -Studien av de kjemiske prosesser i levende organismer, eller sagt på en annen måte; det molekylære grunnlaget
Introduksjon til Biokjemi Ingar Leiros, Institutt for Kjemi, UiT Biokjemi Biokjemi (Wikipedia): -Studien av de kjemiske prosesser i levende organismer, eller sagt på en annen måte; det molekylære grunnlaget
2. GABA A GABA A - GABA GABA GABA B GABA GABA GABA GABA - PRIP- PRIP- GABARAP GABA A PRIP-1 GABA PRIP- PP- PRIP GABARAP PP- Gephyrin
 Folia Pharmacol. Jpn.123 - GABA -PRIP- PRIP- GABARAP PRIP-1 GABA GABA PRIP- PP- PRIP- PP- PRIPGABARAPPP- Gephyrin - -- e-mail: inositol@dent.kyushu-u.ac.jp 1. -GABA GABA B GABA GABA GABA GABA 2. GABA TM
Folia Pharmacol. Jpn.123 - GABA -PRIP- PRIP- GABARAP PRIP-1 GABA GABA PRIP- PP- PRIP- PP- PRIPGABARAPPP- Gephyrin - -- e-mail: inositol@dent.kyushu-u.ac.jp 1. -GABA GABA B GABA GABA GABA GABA 2. GABA TM
Bortezomib (Velcade R, BZ) 26S 3 BZ HDAC6. BZ BZ Bafilomycin A 1. IM-9, RPMI8226, KMS-12-PE LC3B p62 immunoblotting. p62
 Bortezomib (Velcade R,BZ) 2003 FDA 26S 3 BZ BZ BZ Bafilomycin A 1 (ER) Kawaguchi T, Moriya S, et al. Int J Oncol.2011 Clarithromycin(CAM) Azithromycin(AZM) (Nakamura M et al. Int J Oncol. 2010, Renna M,
Bortezomib (Velcade R,BZ) 2003 FDA 26S 3 BZ BZ BZ Bafilomycin A 1 (ER) Kawaguchi T, Moriya S, et al. Int J Oncol.2011 Clarithromycin(CAM) Azithromycin(AZM) (Nakamura M et al. Int J Oncol. 2010, Renna M,
Examination paper for Bi2014 Molecular Biology
 Department of Biology Examination paper for Bi2014 Molecular Biology Academic contact during examination: Professor Atle M. Bones Phone: (+47-)91897237 Examination date/eksamensdag: 8. December 2017 Examination
Department of Biology Examination paper for Bi2014 Molecular Biology Academic contact during examination: Professor Atle M. Bones Phone: (+47-)91897237 Examination date/eksamensdag: 8. December 2017 Examination
NORGES TEKNISK-NATURVITENSKAPELIG UNIVERSITET Side 1 av 5 INSTITUTT FOR FYSIKK. EKSAMEN I FAG CELLEBIOLOGI 1 august 1997 Tid: kl
 NORGES TEKNISK-NATURVITENSKAPELIG UNIVERSITET Side 1 av 5 INSTITUTT FOR FYSIKK Faglig kontakt under eksamen: Navn: Professor Tore Lindmo Tlf.:93432 EKSAMEN I FAG 74618 CELLEBIOLOGI 1 august 1997 Tid: kl
NORGES TEKNISK-NATURVITENSKAPELIG UNIVERSITET Side 1 av 5 INSTITUTT FOR FYSIKK Faglig kontakt under eksamen: Navn: Professor Tore Lindmo Tlf.:93432 EKSAMEN I FAG 74618 CELLEBIOLOGI 1 august 1997 Tid: kl
UNIVERSITETET I OSLO
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Eksamen i MBV 1030 Generell biokjemi Eksamensdag: Mandag 6. desember 2004 Tid for eksamen: kl. 09.00 12.00 Oppgavesettet er på 9 sider Vedlegg:
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Eksamen i MBV 1030 Generell biokjemi Eksamensdag: Mandag 6. desember 2004 Tid for eksamen: kl. 09.00 12.00 Oppgavesettet er på 9 sider Vedlegg:
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet BIOKJEMISK INSTITUTT
 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet BIOKJEMISK INSTITUTT Eksamen i: KJB492 - Bioinformatikk, 3 vekttall Eksamensdag: Onsdag 13.november 2000 Tid for eksamen: kl. 09.00-13.00
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet BIOKJEMISK INSTITUTT Eksamen i: KJB492 - Bioinformatikk, 3 vekttall Eksamensdag: Onsdag 13.november 2000 Tid for eksamen: kl. 09.00-13.00
BI 212- Protein Sorting - Kap. 17 Post-translasjonell modifisering og kvalitetskontroll i r-er (Del 17.6)
 Før de sekretoriske proteiner transporteres videre fra ER-lumen til sitt endelige bestemmelsessted, blir de modifisert ; Dannelse av disulfid-bindinger Foldinger av proteinet Påleiring og prosessering
Før de sekretoriske proteiner transporteres videre fra ER-lumen til sitt endelige bestemmelsessted, blir de modifisert ; Dannelse av disulfid-bindinger Foldinger av proteinet Påleiring og prosessering
NEUROBIOLOGISKE MEKANISMER KNYTTET TIL RUS, AVHENGIGHET OG AVVENNING
 NEUROBIOLOGISKE MEKANISMER KNYTTET TIL RUS, AVHENGIGHET OG AVVENNING Jørg Mørland professor, dr.med. UiO divisjonsdirektør, Folkehelseinstituttet Farmasidagene 2010 28. oktober Det mesolimbiske striocorticale
NEUROBIOLOGISKE MEKANISMER KNYTTET TIL RUS, AVHENGIGHET OG AVVENNING Jørg Mørland professor, dr.med. UiO divisjonsdirektør, Folkehelseinstituttet Farmasidagene 2010 28. oktober Det mesolimbiske striocorticale
EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Mandag 7. mai 2001 Tid: kl Ingen trykte eller håndskrevne hjelpemidler tillatt.
 Side av 5 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK Faglig kontakt under eksamen: Navn: Bjørn Torger Stokke Tlf: 93434 BOKMÅL EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Mandag 7. mai
Side av 5 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK Faglig kontakt under eksamen: Navn: Bjørn Torger Stokke Tlf: 93434 BOKMÅL EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Mandag 7. mai
Supplemental Data Regulation of MBK-2/DYRK by CDK-1 and the Pseudophosphatases EGG-4 and EGG-5 during the Oocyte-to-Embryo Transition
 Cell, Volume 139 Supplemental Data Regulation of MBK-2/DYRK by CDK-1 and the Pseudophosphatases EGG-4 and EGG-5 during the Oocyte-to-Embryo Transition Ken Chih-Chien Cheng, Richard Klancer, Andrew Singson,
Cell, Volume 139 Supplemental Data Regulation of MBK-2/DYRK by CDK-1 and the Pseudophosphatases EGG-4 and EGG-5 during the Oocyte-to-Embryo Transition Ken Chih-Chien Cheng, Richard Klancer, Andrew Singson,
RF Power Capacitors Class1 5kV Discs
 RF Power Capacitors Class 5kV Discs Morgan Advanced Materials is a world leader in the design and manufacture of complex electronic ceramic components and assemblies used in a wide range of applications
RF Power Capacitors Class 5kV Discs Morgan Advanced Materials is a world leader in the design and manufacture of complex electronic ceramic components and assemblies used in a wide range of applications
TRANSKRIPSJONSFAKTORER
 TRANSKRIPSJONSFAKTORER Artikler: *Yang VW. Eukaryotic transkription factors: identification, characterization and functions. J Nutr 1998 Nov 128:11 2045-51 *Chen L. Combinatorialgene regultion by eukaryotic
TRANSKRIPSJONSFAKTORER Artikler: *Yang VW. Eukaryotic transkription factors: identification, characterization and functions. J Nutr 1998 Nov 128:11 2045-51 *Chen L. Combinatorialgene regultion by eukaryotic
Flervalgsoppgaver: proteinsyntese
 Flervalgsoppgaver - proteinsyntese Hver oppgave har ett riktig svaralternativ. Proteinsyntese 1 Hva blir transkribert fra denne DNA sekvensen: 3'-C-C-G-A-A-T-G-T-C-5'? A) 3'-G-G-C-U-U-A-C-A-G-5' B) 3'-G-G-C-T-T-A-C-A-G-5'
Flervalgsoppgaver - proteinsyntese Hver oppgave har ett riktig svaralternativ. Proteinsyntese 1 Hva blir transkribert fra denne DNA sekvensen: 3'-C-C-G-A-A-T-G-T-C-5'? A) 3'-G-G-C-U-U-A-C-A-G-5' B) 3'-G-G-C-T-T-A-C-A-G-5'
europeisk patentskrift
 (12) Oversettelse av europeisk patentskrift (11) NO/EP 2125711 B1 (19) NO NORGE (51) Int Cl. C07C 321/20 (2006.01) A61K 31/216 (2006.01) A61K 31/421 (2006.01) A61K 31/4402 (2006.01) A61K 31/495 (2006.01)
(12) Oversettelse av europeisk patentskrift (11) NO/EP 2125711 B1 (19) NO NORGE (51) Int Cl. C07C 321/20 (2006.01) A61K 31/216 (2006.01) A61K 31/421 (2006.01) A61K 31/4402 (2006.01) A61K 31/495 (2006.01)
RF Power Capacitors Class kV Discs
 RF Power Capacitors Class 0-5kV Discs Morgan Advanced Materials is a world leader in the design and manufacture of complex electronic ceramic components and assemblies used in a wide range of applications
RF Power Capacitors Class 0-5kV Discs Morgan Advanced Materials is a world leader in the design and manufacture of complex electronic ceramic components and assemblies used in a wide range of applications
T h e J o u r n a l o f E x p e r i m e n t a l M e d i c i n e
 T h e J o u r n a l o f E x p e r i m e n t a l M e d i c i n e SUPPLEMENTAL MATERIAL Nishikawa et al., http://www.jem.org/cgi/content/full/jem.20092144/dc1 JEM S1 Figure S1. Flow cytometric analysis of
T h e J o u r n a l o f E x p e r i m e n t a l M e d i c i n e SUPPLEMENTAL MATERIAL Nishikawa et al., http://www.jem.org/cgi/content/full/jem.20092144/dc1 JEM S1 Figure S1. Flow cytometric analysis of
4. Antigenpresentasjon til T celler. MHC molekyler.
 Immunologi 3. semester, V2015 4. Antigenpresentasjon til T celler. MHC molekyler. Karl Schenck Institutt for oral biologi TcR og MHC molekyler Noen hovedgrupper T celler som forlater thymus T celler med
Immunologi 3. semester, V2015 4. Antigenpresentasjon til T celler. MHC molekyler. Karl Schenck Institutt for oral biologi TcR og MHC molekyler Noen hovedgrupper T celler som forlater thymus T celler med
Regularitati ascunse si corelatii in nano-bio-structuri
 NATIONAL INSTITUTE OF MATERIALS PHYSICS BUCHAREST-MAGURELE Atomistilor Str. bis, P.O. Box MG-7, 77 Magurele-Ilfov, Romania Phone: +() 98, Fax: +() 977, email: pintilie@infim.ro, http://www.infim.ro Regularitati
NATIONAL INSTITUTE OF MATERIALS PHYSICS BUCHAREST-MAGURELE Atomistilor Str. bis, P.O. Box MG-7, 77 Magurele-Ilfov, Romania Phone: +() 98, Fax: +() 977, email: pintilie@infim.ro, http://www.infim.ro Regularitati
Besvarelse SIF4070 Cellebiologi 31. mai 2002
 1 Besvarelse SIF4070 Cellebiologi 31. mai 2002 Oppgave 1: Syntese av plasmamembranen. Transport over plasmamembranen a) Proteinet når ER Transport til ER membranen Transmembranproteiner som skal til ER
1 Besvarelse SIF4070 Cellebiologi 31. mai 2002 Oppgave 1: Syntese av plasmamembranen. Transport over plasmamembranen a) Proteinet når ER Transport til ER membranen Transmembranproteiner som skal til ER
Compounding Altered- Release Pharmaceuticals
 Slide 1 Compounding Altered- Release Pharmaceuticals Part IV Loyd V. Allen, Jr., Ph.D. Slide 2 Compounding Altered-Release Pharmaceuticals-Part Part IV Parenteral Miscellaneous Drug Examples that are Good
Slide 1 Compounding Altered- Release Pharmaceuticals Part IV Loyd V. Allen, Jr., Ph.D. Slide 2 Compounding Altered-Release Pharmaceuticals-Part Part IV Parenteral Miscellaneous Drug Examples that are Good
EKSAMEN I EMNE SIF4070 CELLEBIOLOGI
 Side 1 av 7 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Faglig kontakt under eksamen: professor Catharina Davies Tel 73593688 eller 41666231
Side 1 av 7 NORGES TEKNISK-NATURVITENSKAPELIGE UNIVERSITET INSTITUTT FOR FYSIKK EKSAMEN I EMNE SIF4070 CELLEBIOLOGI Faglig kontakt under eksamen: professor Catharina Davies Tel 73593688 eller 41666231
Sammenligningen mellom Arabidopsis thaliana genomet og de kjente genomene fra cyanobakterier, gjær, bananflue og nematode, viser bl. a.
 Sammenligningen mellom Arabidopsis thaliana genomet og de kjente genomene fra cyanobakterier, gjær, bananflue og nematode, viser bl. a. Antall gener som er involvert i cellulær kommunikasjon og signaloverføring
Sammenligningen mellom Arabidopsis thaliana genomet og de kjente genomene fra cyanobakterier, gjær, bananflue og nematode, viser bl. a. Antall gener som er involvert i cellulær kommunikasjon og signaloverføring
Søkers navn Søknadstype Institusjon Prosjekttittel Sum
 Anita Ryningen Post-transplant adoptive t-cell immunotherapy Anne Margrete Øyan Identifying new therapeutic compounds against cancer using in vitro and in vivo screening models - basic mechanisms and developments
Anita Ryningen Post-transplant adoptive t-cell immunotherapy Anne Margrete Øyan Identifying new therapeutic compounds against cancer using in vitro and in vivo screening models - basic mechanisms and developments
Andrew Gendreau, Olga Rosenbaum, Anthony Taylor, Kenneth Wong, Karl Dusen
 Andrew Gendreau, Olga Rosenbaum, Anthony Taylor, Kenneth Wong, Karl Dusen The Process Goal Definition Data Collection Data Preprocessing EDA Choice of Variables Choice of Method(s) Performance Evaluation
Andrew Gendreau, Olga Rosenbaum, Anthony Taylor, Kenneth Wong, Karl Dusen The Process Goal Definition Data Collection Data Preprocessing EDA Choice of Variables Choice of Method(s) Performance Evaluation
EKSAMENSOPPGAVE I MOL4010 Molekylærbiologi for teknologer
 Fag/ mnekode: MOL4010 Dato: 04.06.2010 Studentnr.: 1 Det medisinske fakultet Institutt for kreftforskning og molekylær medisin KSAMNSOPPGAV I MOL4010 Molekylærbiologi for teknologer Fredag 4. juni 2010,
Fag/ mnekode: MOL4010 Dato: 04.06.2010 Studentnr.: 1 Det medisinske fakultet Institutt for kreftforskning og molekylær medisin KSAMNSOPPGAV I MOL4010 Molekylærbiologi for teknologer Fredag 4. juni 2010,
OECD GUIDELINE FOR THE TESTING OF CHEMICALS
 TG 442C OECD GUIDELINE FOR THE TESTING OF CHEMICALS In Chemico Skin Sensitisation: Direct Peptide Reactivity Assay (DPRA) INTRODUCTION in chemico in chemicoin vitro in silico in chemico in vitro OECD,
TG 442C OECD GUIDELINE FOR THE TESTING OF CHEMICALS In Chemico Skin Sensitisation: Direct Peptide Reactivity Assay (DPRA) INTRODUCTION in chemico in chemicoin vitro in silico in chemico in vitro OECD,
